相关产品推荐更多 >
万千商家帮你免费找货
0 人在求购买到急需产品
- 详细信息
- 询价记录
- 文献和实验
- 技术资料
- 库存:
999
- 英文名:
RapiDxFire Thermostable Reverse Transcriptase
- 供应商:
中北林格
- 规格:
50 rxns
- 在高温下极其活跃(55-80°C):通过各种RNA模板提高cDNA合成的特异性
- 敏感: 在两步RT-qPCR分析中检测≤100拷贝的RNA
- 反应时间短(5分钟或更短): 简化的RT-qPCR工作流程和更快的出结果时间
- 在室温下稳定(> 3个月):简化自动化卡座上的设置(无需冷藏),可在冷藏有限的环境中使用
- Lyo兼容: 酶制剂不含甘油和其他成分,不会干扰下游冻干步骤
- 批次间重复性:在ISO 13485认证的设施中制造。
酶包装大小(50或250 Rxns)基于25μL反应体积。
RapiDxFire热稳定逆转录酶是一种显着优于流行的MMLV和AMV逆转录酶的方法。最初在温泉中发现,这种强大的噬菌体酶在升高的温度下表现出越来越高的活性(在90℃下10分钟后活性保持约60%)并且在20-25℃下储存> 3个月后保持其核心活性。这种酶缺乏RNAse H和3'→5'核酸外切酶活性并有效合成短cDNA片段(≤1Kb)。与其他逆转录酶一样,RapiDxFire RT具有DNA聚合酶活性,但缺乏5'→3'核酸外切酶活性。酶的储存缓冲液不含甘油和Triton™X-100,可进一步优化用于下游冻干步骤。
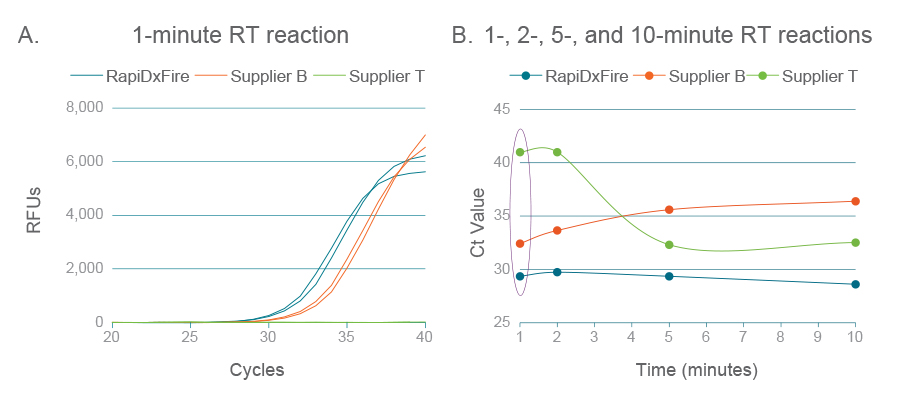
RapiDxFire耐高温RT在高温下表现最佳

3个月室温稳定性

产品名称: RapiDxFire Thermostable ReverseTranscriptase, 3Units/µL
规格:50次, 250次
对应货号: 30250-1 30250-2
欢迎来电咨询订购: 010-88891086 www.bjzblg.com QQ:1443274331
风险提示:丁香通仅作为第三方平台,为商家信息发布提供平台空间。用户咨询产品时请注意保护个人信息及财产安全,合理判断,谨慎选购商品,商家和用户对交易行为负责。对于医疗器械类产品,请先查证核实企业经营资质和医疗器械产品注册证情况。
- 作者
- 内容
- 询问日期
 文献和实验
文献和实验I think the real secret here is the removal of secondary structure by the thermostable RT coupled with the immobilized RNA forming a "reusable" pool of template. This works well with total RNA or a poly(A)+ fraction.
here is the removal of secondary structure by the thermostable RT coupled with the immobilized RNA forming a "reusable" pool of template. This works well with total RNA or a poly(A)+ fraction.
of secondary structure by the thermostable RT coupled with the immobilized RNA forming a "reusable" pool of template. This works well with total RNA or a poly(A) fraction. 生物试剂库: 试剂库 :http://www.bioon.com.cn/reagent












