产品简介:
该试剂盒用于构建高质量的二代测序文库(illumina和MGI平台)。该试剂盒需要双链DNA片段(钝性和/或粘性)作为NGS建库的输入DNA,并与酶法和物理方法(超声、雾化等)产生的DNA片段兼容。不同类型的index可以实现库的多路复用。
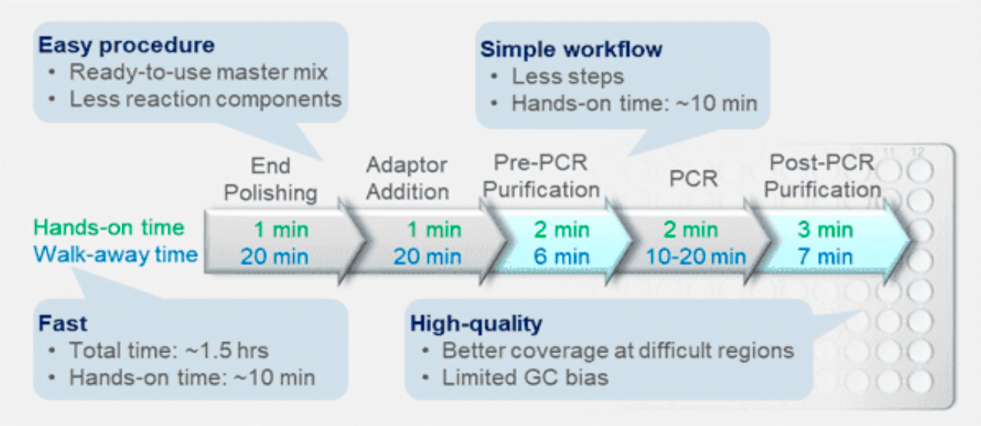
优化后的试剂盒,用于不同类型样本的二代测序(NGS)建库。大多数DNA建库需要将剪切后的DNA片段连接到文库适配器上,而建库与NGS数据的质量密切相关。BioDynami凭借独特的DNA文库制备技术,NGS文库制备时间约为1.5h,人工操作时间仅需10分钟。
The kit was developed for construction of high quality libraries for next generation sequencing (illumina and MGI Platforms). The kit needs double strand DNA fragments (blunt and/or sticky) as input DNA for NGS library preparation, and is compatible with DNA fragments generated from both enzymatic methods and physical methods (sonication, nebulization etc.). Library multiplexing is possible with different types of indexes.
The kit was optimized for next generation sequencing (NGS) library preparation with different types of samples. Most of DNA library preparation requires the ligation of sheared DNA fragments to library adaptors and the DNA library preparation is closely related to the quality of NGS data. With BioDynami’s unique DNA library preparation technologies, the fast and simple kit allows high quality NGS library preparation to be completed in 1.5 hours with only 10 minutes of hands-on time.

一些基因组区域很难被均匀覆盖,通常导致这些区域的覆盖率很低或存在差距。
典型的难点:
★ GC含量高;
★ 有二级结构:主要是由于重复序列;
★ 最坏的情况:同时具有高GC含量和重复序列。
如上图所示,人类TERT基因是最困难的区域之一。NGS数据显示,BioDynami试剂盒在覆盖极难识别的人类TERT基因区域方面表现最好。
Some genomic regions are very difficult to be covered evenly and usually result in very low coverage rate or gap in these regions.
Typical difficult regions are:
• with high GC contents
• have secondary structures: mainly due to repeat sequences
• the worst cases: have both high GC contents and repeated sequences.
Example: human TERT gene is one of the most difficult regions as shown above. NGS data showed that BioDynami kit has the best performance to cover the extremely difficult human TERT gene region.

illumina试剂盒有三种index类型:
Non-index(illumina Cat.# 30009):文库没有索引。
Index(illumina Cat.# 30021):每个引物都包含6个碱基的独特条形码序列。RNA测序文库复用可用于多达48个样本。
Unique dual index(illumina Cat.# 30023):使用独特的双索引,可实现多达96个样本的RNA测序文库复用。建立了4-Base Difference Index System。该系统可以在8个碱基索引区域生成至少4个不同于其他碱基的索引。独特的双索引引物可识别测序错误,如索引跳变、分配错误和解复用错误。
Three index types are available for the illumina platform kits:
Non-index (illumina Cat.# 30009): Libraries do not have index.
Index (illumina Cat.# 30021): A unique barcode sequence with 6 bases has been included in each of the index primers. RNA Sequencing library multiplexing is possible with up to 48 samples.
Unique dual index (illumina Cat.# 30023): RNA Sequencing library multiplexing up to 96 samples is possible with the unique dual indexes. We have developed a 4-Base Difference Index System. The system can generate indexes with at least 4 bases different from others in the 8-base indexing region. the unique dual indexing primers identify sequencing errors such as index hopping, mis-assignment, and de-multiplexing errors.
Indexes are available for the MGI platform kits (Cat.# 34021).
产品订购:
| 产品名称 |
货号 |
产品规格 |
| NGS DNA Library Prep Kit (illumina) |
30009S |
non-index,24 reactions |
| NGS DNA Library Prep Kit (illumina) |
30009L |
non-index,48 reactions |
| NGS DNA Library Prep Kit (illumina) |
30021S |
index,24 reactions |
| NGS DNA Library Prep Kit (illumina) |
30021L |
index,48 reactions |
| NGS DNA Library Prep Kit (illumina) |
30023S |
unique dual index,96 reactions |
| NGS DNA Library Prep Kit (illumina) |
30023L |
unique dual index,192 reactions |
| NGS DNA Library Prep Kit(MGI) |
34021S |
index,24 reactions |
| NGS DNA Library Prep Kit(MGI) |
34021L |
index,48 reactions |
