| 出品公司: | |
|---|---|
| 别名: | pET41b, pET 41b |
| 质粒类型: | 大肠杆菌蛋白表达 |
| 表达水平: | 高 |
| 克隆方法: | 多克隆位点,限制性内切酶 |
| 载体大小: | 5932 bp |
| 5' 测序引物: | T7或pGEX-5‘ |
| 5' 测序引物序列: | T7: 5'-TAATACGACTCACTATAGGG-3'; pGEX-5': 5'-GGGCTGGCAAGCCACGTTTGGTG-3' |
| 载体标签: | N-GST, N-His, N-Thrombin |
| 载体抗性: | Kanamycin |
| 表达宿主菌: | BL21(DE3) 、 Rosetta2(DE3)、C43(DE3) |
| 备注: |
Encodes GST fusion tag; Nterm thrombin cleavage site; Nterm enterokinase cleavage site; PshAI blunt cloning site; a,b,c vary by MCS
|
| 产品目录号: | ZY70557-3 |
| 稳定性: | 瞬时表达 Transient |
| 组成型: | 组成型 Constitutive |
| 病毒/非病毒: | 非病毒 |
万千商家帮你免费找货
0 人在求购买到急需产品
- 详细信息
- 文献和实验
- 技术资料
- 保存条件:
-20℃低温保存
- 保质期:
三年
- 英文名:
pET-41b(+)
- 库存:
20
- 供应商:
泽叶生物
pET41b载体基本信息
pET41b载体质粒图谱和多克隆位点信息
| Feature Name | Start | End | |
|---|---|---|---|
| T7_terminator | 120 | 1 | |
| T7_Terminal_primer | 69 | 87 | |
| EK | 278 | 264 | |
| S15 | 353 | 309 | |
| 6xHIS | 413 | 396 | |
| GST (variant) | 1094 | 444 | |
| T7_transl_en_RBS | 1119 | 1103 | |
| lacO | 1164 | 1137 | |
| T7_promoter | 1182 | 1164 | |
| tet (300 - 563) | 1218 | 1481 | |
| pBRrevBam_primer | 1289 | 1270 | |
| lacI | 1564 | 2655 | |
| ROP | 3227 | 3418 | |
| pGEX_3_primer | 3434 | 3412 | |
| pBR322_origin | 4452 | 3833 | |
| KanR2 | 4558 | 5373 |
| ORF | Start | End | |
|---|---|---|---|
| ORF frame 3 | 1094 | 147 | |
| ORF frame 3 | 1095 | 1823 | |
| ORF frame 1 | 1696 | 2655 | |
| ORF frame 1 | 4558 | 5373 |
| Enzyme Name | Cut |
|---|---|
| XhoI | 174 |
| NotI | 182 |
| EagI | 182 |
| HindIII | 189 |
| SalI | 195 |
| SacI | 206 |
| PstI | 214 |
| AscI | 216 |
| StuI | 226 |
| EcoRI | 235 |
| BamHI | 241 |
| EcoRV | 251 |
| NcoI | 254 |
| AgeI | 291 |
| KpnI | 298 |
| BglII | 301 |
| SacII | 392 |
| SpeI | 423 |
| MscI | 886 |
| NdeI | 1093 |
| XbaI | 1131 |
| ApaI | 2134 |
| HpaI | 2429 |
| NruI | 4646 |
| SmaI | 4863 |
| XmaI | 4861 |
pET41b载体简介
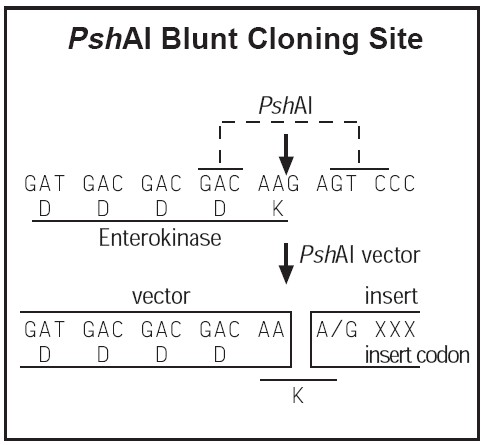
The pET-41 series is designed for cloning and high-level expression of peptide sequences fused with the 220 aa GST•Tag™ protein. Unique sites are shown on the circle map. Note that the sequence is numbered by the pBR322 convention, so the T7 expression region is reversed on the circle map. The cloning/expression region of the coding strand transcribed by T7 RNA polymerase is shown below. The f1 origin is oriented so that infection with helper phage will produce virions containing single stranded DNA that corresponds to the coding strand. Therefore, single stranded sequencing should be performed using the T7 terminator primer (cat. no. 69337-3).Vector encoded sequence can be completely removed when cloning into the PshAI site (as shown below) and then cleaving the GST fusion protein with Enterokinase.
pET41b载体序列
DEFINITION pET-41 b(+)
ACCESSION
KEYWORDS
SOURCE
ORGANISM other sequences; artificial sequences; vectors.
COMMENT This file is created by Vector NTI
http://www.biofeng.com/
COMMENT ORIGDB|GenBank
COMMENT VNTAUTHORNAME|biofeng.com|
COMMENT VNTOAUTHORAD1|Novagen Inc.|
FEATURES Location/Qualifiers
source 1..5932
/organism="pET-41 b(+)"
/mol_type="other DNA"
terminator complement(1..120)
/label="T7_terminator"
misc_feature 69..87
/label="T7_Terminal_primer"
CDS complement(147..1094)
/label="ORF frame 3"
/translation="MSPILGYWKIKGLVQPTRLLLEYLEEKYEEHLYERDEGDKWRNK
KFELGLEFPNLPYYIDGDVKLTQSMAIIRYIADKHNMLGGCPKERAEISMLEGAVLDI
RYGVSRIAYSKDFETLKVDFLSKLPEMLKMFEDRLCHKTYLNGDHVTHPDFMLYDALD
VVLYMDPMCLDAFPKLVCFKKRIEAIPQIDKYLKSSKYIAWPLQGWQATFGGGDHPPK
SDGSTSGSGHHHHHHSAGLVPRGSTAIGMKETAAAKFERQHMDSPDLGTGGGSGDDDD
KSPMDIGDPNSVQALARLQASSVDKLAAALEHHHHHHHH*"
misc_feature complement(264..278)
/label="EK"
/translation="MSPILGYWKIKGLVQPTRLLLEYLEEKYEEHLYERDEGDKWRNK
KFELGLEFPNLPYYIDGDVKLTQSMAIIRYIADKHNMLGGCPKERAEISMLEGAVLDI
RYGVSRIAYSKDFETLKVDFLSKLPEMLKMFEDRLCHKTYLNGDHVTHPDFMLYDALD
VVLYMDPMCLDAFPKLVCFKKRIEAIPQIDKYLKSSKYIAWPLQGWQATFGGGDHPPK
SDGSTSGSGHHHHHHSAGLVPRGSTAIGMKETAAAKFERQHMDSPDLGTGGGSGDDDD
KSPMDIGDPNSVQALARLQASSVDKLAAALEHHHHHHHH*"
misc_feature complement(309..353)
/label="S15"
/translation="MSPILGYWKIKGLVQPTRLLLEYLEEKYEEHLYERDEGDKWRNK
KFELGLEFPNLPYYIDGDVKLTQSMAIIRYIADKHNMLGGCPKERAEISMLEGAVLDI
RYGVSRIAYSKDFETLKVDFLSKLPEMLKMFEDRLCHKTYLNGDHVTHPDFMLYDALD
VVLYMDPMCLDAFPKLVCFKKRIEAIPQIDKYLKSSKYIAWPLQGWQATFGGGDHPPK
SDGSTSGSGHHHHHHSAGLVPRGSTAIGMKETAAAKFERQHMDSPDLGTGGGSGDDDD
KSPMDIGDPNSVQALARLQASSVDKLAAALEHHHHHHHH*"
misc_feature complement(396..413)
/label="6xHIS"
/translation="MSPILGYWKIKGLVQPTRLLLEYLEEKYEEHLYERDEGDKWRNK
KFELGLEFPNLPYYIDGDVKLTQSMAIIRYIADKHNMLGGCPKERAEISMLEGAVLDI
RYGVSRIAYSKDFETLKVDFLSKLPEMLKMFEDRLCHKTYLNGDHVTHPDFMLYDALD
VVLYMDPMCLDAFPKLVCFKKRIEAIPQIDKYLKSSKYIAWPLQGWQATFGGGDHPPK
SDGSTSGSGHHHHHHSAGLVPRGSTAIGMKETAAAKFERQHMDSPDLGTGGGSGDDDD
KSPMDIGDPNSVQALARLQASSVDKLAAALEHHHHHHHH*"
gene complement(444..1094)
/label="GST (variant)"
/gene="GST (variant)"
/translation="MSPILGYWKIKGLVQPTRLLLEYLEEKYEEHLYERDEGDKWRNK
KFELGLEFPNLPYYIDGDVKLTQSMAIIRYIADKHNMLGGCPKERAEISMLEGAVLDI
RYGVSRIAYSKDFETLKVDFLSKLPEMLKMFEDRLCHKTYLNGDHVTHPDFMLYDALD
VVLYMDPMCLDAFPKLVCFKKRIEAIPQIDKYLKSSKYIAWPLQGWQATFGGGDHPPK
SDGSTSGSGHHHHHHSAGLVPRGSTAIGMKETAAAKFERQHMDSPDLGTGGGSGDDDD
KSPMDIGDPNSVQALARLQASSVDKLAAALEHHHHHHHH*"
CDS 1095..1823
/label="ORF frame 3"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
misc_feature complement(1103..1119)
/label="T7_transl_en_RBS"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
misc_feature complement(1137..1164)
/label="lacO"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
promoter complement(1164..1182)
/label="T7_promoter"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
gene 1218..1481
/label="tet (300 - 563)"
/gene="tet (300 - 563)"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
misc_feature complement(1270..1289)
/label="pBRrevBam_primer"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
misc_feature 1564..2655
/label="lacI"
/translation="MYISFLKLNKIISRGELLSAHNSPIVSRINFAGSRSISILYAGR
IVAGITGATGAVAGAYIADITDGEDRARHFGLMSACFGVGMVAGPVAGGLLGAISLHA
PFLAAAVLNGLNLLLGCFLMQESHKGERRDPGHHRMAQNLSRYGMIAPGRESIQGGEC
ETSNVIRCRRVCRCLLSDRFPRGEPGQPRFCENAGKSGSGDGGAELHSQPRGTTTGGQ
TVVADWRCHLQSGPARAVANCRGD*"
CDS 1696..2655
/label="ORF frame 1"
/translation="MAELNYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKS
RADQLGASVVVSMVERSGVEACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNV
PALFLDVSDQTPINSIIFSHEDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGW
HKYLTRNQIQPIAEREGDWSAMSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITE
SGLRVGADISVVGYDDTEDSSCYIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQ
LLPVSLVKRKTTLAPNTQTASPRALADSLMQLARQVSRLESGQ*"
misc_feature 3227..3418
/label="ROP"
/translation="MAELNYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKS
RADQLGASVVVSMVERSGVEACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNV
PALFLDVSDQTPINSIIFSHEDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGW
HKYLTRNQIQPIAEREGDWSAMSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITE
SGLRVGADISVVGYDDTEDSSCYIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQ
LLPVSLVKRKTTLAPNTQTASPRALADSLMQLARQVSRLESGQ*"
misc_feature complement(3412..3434)
/label="pGEX_3_primer"
/translation="MAELNYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKS
RADQLGASVVVSMVERSGVEACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNV
PALFLDVSDQTPINSIIFSHEDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGW
HKYLTRNQIQPIAEREGDWSAMSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITE
SGLRVGADISVVGYDDTEDSSCYIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQ
LLPVSLVKRKTTLAPNTQTASPRALADSLMQLARQVSRLESGQ*"
rep_origin complement(3833..4452)
/label="pBR322_origin"
/translation="MAELNYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKS
RADQLGASVVVSMVERSGVEACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNV
PALFLDVSDQTPINSIIFSHEDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGW
HKYLTRNQIQPIAEREGDWSAMSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITE
SGLRVGADISVVGYDDTEDSSCYIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQ
LLPVSLVKRKTTLAPNTQTASPRALADSLMQLARQVSRLESGQ*"
gene 4558..5373
/label="KanR2"
/gene="KanR2"
/translation="MAELNYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKS
RADQLGASVVVSMVERSGVEACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNV
PALFLDVSDQTPINSIIFSHEDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGW
HKYLTRNQIQPIAEREGDWSAMSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITE
SGLRVGADISVVGYDDTEDSSCYIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQ
LLPVSLVKRKTTLAPNTQTASPRALADSLMQLARQVSRLESGQ*"
CDS 4558..5373
/label="ORF frame 1"
/translation="MSHIQRETSCSRPRLNSNMDADLYGYKWARDNVGQSGATIYRLY
GKPDAPELFLKHGKGSVANDVTDEMVRLNWLTEFMPLPTIKHFIRTPDDAWLLTTAIP
GKTAFQVLEEYPDSGENIVDALAVFLRRLHSIPVCNCPFNSDRVFRLAQAQSRMNNGL
VDASDFDDERNGWPVEQVWKEMHKLLPFSPDSVVTHGDFSLDNLIFDEGKLIGCIDVG
RVGIADRYQDLAILWNCLGEFSPSLQKRLFQKYGIDNPDMNKLQFHLMLDEFF*"
ORIGIN
1 atccggatat agttcctcct ttcagcaaaa aacccctcaa gacccgttta gaggccccaa
61 ggggttatgc tagttattgc tcagcggtgg cagcagccaa ctcagcttcc tttcgggctt
121 tgtttagcag cctaggtatt aatcaattag tggtggtggt ggtggtggtg gtgctcgagt
181 gcggccgcaa gcttgtcgac ggagctcgcc tgcaggcgcg ccaaggcctg tacagaattc
241 ggatccccga tatccatggg actcttgtcg tcgtcatcac cggagccacc accggtaccc
301 agatctgggc tgtccatgtg ctggcgttcg aatttagcag cagcggtttc tttcatacca
361 attgcagtac taccgcgtgg caccagaccc gcggagtgat ggtgatggtg atgaccagaa
421 ccactagttg aaccatccga ttttggagga tggtcgccac caccaaacgt ggcttgccag
481 ccctgcaaag gccatgctat atacttgctg gatttcaagt acttatcaat ttgtgggata
541 gcttcaatac gttttttaaa acaaactaat tttgggaacg catccaggca cattgggtcc
601 atgtataaaa caacatcaag agcgtcatac aacatgaagt caggatgggt tacatgatca
661 ccatttaaat atgttttatg acataaacga tcttcgaaca ttttcagcat ttcaggtagc
721 ttgctaagaa aatcaacttt gagagtttca aagtctttac tatatgcaat tctcgaaaca
781 ccgtatctaa tatccaaaac cgctccttca agcattgaaa tctctgcacg ctcttttgga
841 caaccaccca acatgttgtg cttgtcagct atataacgta tgatggccat agactgtgtt
901 aatttaacat caccatcaat ataataagga agattgggaa actccaaacc caattcaaac
961 tttttgtttc gccatttatc accttcatcg cgctcataca aatgctcttc atatttttct
1021 tcaagatatt ccaaaagaag tcgagtgggt tgcacaaggc ccttaatttt ccaataacct
1081 agtatagggg acatatgtat atctccttct taaagttaaa caaaattatt tctagagggg
1141 aattgttatc cgctcacaat tcccctatag tgagtcgtat taatttcgcg ggatcgagat
1201 cgatctcgat cctctacgcc ggacgcatcg tggccggcat caccggcgcc acaggtgcgg
1261 ttgctggcgc ctatatcgcc gacatcaccg atggggaaga tcgggctcgc cacttcgggc
1321 tcatgagcgc ttgtttcggc gtgggtatgg tggcaggccc cgtggccggg ggactgttgg
1381 gcgccatctc cttgcatgca ccattccttg cggcggcggt gctcaacggc ctcaacctac
1441 tactgggctg cttcctaatg caggagtcgc ataagggaga gcgtcgagat cccggacacc
1501 atcgaatggc gcaaaacctt tcgcggtatg gcatgatagc gcccggaaga gagtcaattc
1561 agggtggtga atgtgaaacc agtaacgtta tacgatgtcg cagagtatgc cggtgtctct
1621 tatcagaccg tttcccgcgt ggtgaaccag gccagccacg tttctgcgaa aacgcgggaa
1681 aaagtggaag cggcgatggc ggagctgaat tacattccca accgcgtggc acaacaactg
1741 gcgggcaaac agtcgttgct gattggcgtt gccacctcca gtctggccct gcacgcgccg
1801 tcgcaaattg tcgcggcgat taaatctcgc gccgatcaac tgggtgccag cgtggtggtg
1861 tcgatggtag aacgaagcgg cgtcgaagcc tgtaaagcgg cggtgcacaa tcttctcgcg
1921 caacgcgtca gtgggctgat cattaactat ccgctggatg accaggatgc cattgctgtg
1981 gaagctgcct gcactaatgt tccggcgtta tttcttgatg tctctgacca gacacccatc
2041 aacagtatta ttttctccca tgaagacggt acgcgactgg gcgtggagca tctggtcgca
2101 ttgggtcacc agcaaatcgc gctgttagcg ggcccattaa gttctgtctc ggcgcgtctg
2161 cgtctggctg gctggcataa atatctcact cgcaatcaaa ttcagccgat agcggaacgg
2221 gaaggcgact ggagtgccat gtccggtttt caacaaacca tgcaaatgct gaatgagggc
2281 atcgttccca ctgcgatgct ggttgccaac gatcagatgg cgctgggcgc aatgcgcgcc
2341 attaccgagt ccgggctgcg cgttggtgcg gacatctcgg tagtgggata cgacgatacc
2401 gaagacagct catgttatat cccgccgtta accaccatca aacaggattt tcgcctgctg
2461 gggcaaacca gcgtggaccg cttgctgcaa ctctctcagg gccaggcggt gaagggcaat
2521 cagctgttgc ccgtctcact ggtgaaaaga aaaaccaccc tggcgcccaa tacgcaaacc
2581 gcctctcccc gcgcgttggc cgattcatta atgcagctgg cacgacaggt ttcccgactg
2641 gaaagcgggc agtgagcgca acgcaattaa tgtaagttag ctcactcatt aggcaccggg
2701 atctcgaccg atgcccttga gagccttcaa cccagtcagc tccttccggt gggcgcgggg
2761 catgactagc atgatcgtgc tcctgtcgtt gaggacccgg ctaggctggc ggggttgcct
2821 tactggttag cagaatgaat caccgatacg cgagcgaacg tgaagcgact gctgctgcaa
2881 aacgtctgcg acctgagcaa caacatgaat ggtcttcggt ttccgtgttt cgtaaagtct
2941 ggaaacgcgg aagtcagcgc cctgcaccat tatgttccgg atctgcatcg caggatgctg
3001 ctggctaccc tgtggaacac ctacatctgt attaacgaag cgctggcatt gaccctgagt
3061 gatttttctc tggtcccgcc gcatccatac cgccagttgt ttaccctcac aacgttccag
3121 taaccgggca tgttcatcat cagtaacccg tatcgtgagc atcctctctc gtttcatcgg
3181 tatcattacc cccatgaaca gaaatccccc ttacacggag gcatcagtga ccaaacagga
3241 aaaaaccgcc cttaacatgg cccgctttat cagaagccag acattaacgc ttctggagaa
3301 actcaacgag ctggacgcgg atgaacaggc agacatctgt gaatcgcttc acgaccacgc
3361 tgatgagctt taccgcagct gcctcgcgcg tttcggtgat gacggtgaaa acctctgaca
3421 catgcagctc ccggagacgg tcacagcttg tctgtaagcg gatgccggga gcagacaagc
3481 ccgtcagggc gcgtcagcgg gtgttggcgg gtgtcggggc gcagccatga cccagtcacg
3541 tagcgatagc ggagtgtata ctggcttaac tatgcggcat cagagcagat tgtactgaga
3601 gtgcaccata tatgcggtgt gaaataccgc acagatgcgt aaggagaaaa taccgcatca
风险提示:丁香通仅作为第三方平台,为商家信息发布提供平台空间。用户咨询产品时请注意保护个人信息及财产安全,合理判断,谨慎选购商品,商家和用户对交易行为负责。对于医疗器械类产品,请先查证核实企业经营资质和医疗器械产品注册证情况。
 文献和实验
文献和实验表达载体的构建是生物技术和所需蛋白质产生的基本工具,本文总结了pET-32α(+)载体技术的构建,该技术通常用于研究实验室。该方法包括获得用于构建载体的外源DNA片段,将DNA片段亚克隆到pET-32α(+)表达载体中,在大肠杆菌 BL21(DE3)中进行蛋白质表达以及在大肠杆菌中在天然条件下进行蛋白质纯化。裂解液。介绍作为分子生物学实验室中非常有用的实践,重组DNA技术(或基因克隆)是指将DNA片段从一种生物体转移到自我复制的遗传元件如表达载体。然后插入的DNA可以在外源宿主细胞中繁殖
相关专题 1)噬菌体DE3溶源化的菌株如BL21(DE3)是最常用的表达菌株,噬菌体DE3是λ噬菌体的衍生株,构建好的表达载体 可以直接转入表达菌株中,诱导蛋白的表达方式与lac启动子一样都是IPTG诱导。 2)表达菌株BL21(DE3)上带有lacI基因,T7 RNA 聚合酶编码基因以及lacUV5启动子,lacUV5启动子可以启动T7 RNA 聚合酶的表达; 3)表达质粒 如pET-28质粒带有T7启动子和编码阻遏
( DE3 )表达的活性蛋白比其它宿主菌高 10 倍以上。 Origami 宿主菌与氨苄抗性质粒相容,可用于 pET-32 载体,硫氧还蛋白标签能够进一步增强在胞浆内形成二硫键。 TrxB 和 gor 突变可分别用卡那霉素和四环素选择,因此该菌株建议用于带氨苄抗性标记 bla 的 pET 质粒。 Origami B 宿主菌来源于 BL21 lacZY 突变株,还带有与原始 Origami 菌株相同的 TrxB / gor 突变。 Origami B 菌株集 BL21 、 Tuner
 技术资料
技术资料暂无技术资料 索取技术资料