手机验证

/
HEP-G2; Hep G2; HEP G2; HepG2; HEPG2;Hep-G2
/
充足
/
/
/
/
/
/
人肝癌细胞
人
/
上皮细胞样
/
/
复苏发货或干冰冷冻发货
/
贴壁
1 x 10^6 cells/vial/T25
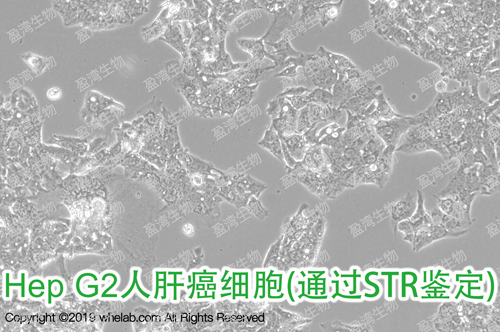
细胞名称:Hep G2人肝癌细胞(STR鉴定正确)
产品货号:C1110
培养条件:MEM+10%FBS+1%PS (配套完全培养基:M0301)
生长特性:贴壁
细胞形态:上皮样
产品规格:1 x 10^6 cells/vial/T25
| 产品货号 | 细胞名称 | 中文名称 | 生长特性 | 细胞形态 | 产品规格 |
| C1010 | HCCLM3 | 人肝癌细胞( 高转) | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1011 | Hep 3B2.1-7 | 人肝癌细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1013 | HuH-7 | 人肝癌细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1027 | PLC/PRF/5 | 人肝癌亚力山 大细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1030 | BEL-7404 | 人肝癌细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1035 | MHCC97-L | 人肝癌细胞( 低转) | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1072 | HCCC-9810 | 人胆管细胞型 肝癌细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1093 | Li-7 | 人肝癌细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1117 | MHCC97-H | 人肝癌细胞( 高转) | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1193 | SK-HEP-1 | 人肝癌细胞 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
| C1322 | SMMC7721/DDP | 人肝癌顺铂耐 药株 | 贴壁 | 上皮样 | 1 x 10^6 cells/vial/T25 |
细胞在发货前进行质检:
1、细胞存种的活性检测;
2、细菌、真菌、霉菌污染物镜检;
3、衣原体、支原体检测。


联系人:胡经理
电话:15921028707(微信同号) :3407506161
:3407506161 :021-55150076
:021-55150076
官网:https://whelab.com
公司邮箱:info@whelab.com
技术邮箱:tech@whelab.com
地址:上海宝山区呼兰西路129号8号楼4层
风险提示:丁香通仅作为第三方平台,为商家信息发布提供平台空间。用户咨询产品时请注意保护个人信息及财产安全,合理判断,谨慎选购商品,商家和用户对交易行为负责。对于医疗器械类产品,请先查证核实企业经营资质和医疗器械产品注册证情况。
 询价记录
询价记录
 文献和实验
文献和实验
PubMed=233137; DOI=10.1038/282615a0
Aden D.P., Fogel A., Plotkin S.A., Damjanov I., Knowles B.B.
Controlled synthesis of HBsAg in a differentiated human liver carcinoma-derived cell line.
Nature 282:615-616(1979)
PubMed=6248960; DOI=10.1126/science.6248960
Knowles B.B., Howe C.C., Aden D.P.
Human hepatocellular carcinoma cell lines secrete the major plasma proteins and hepatitis B surface antigen.
Science 209:497-499(1980)
PubMed=6288577; DOI=10.1002/ijc.2910300106
Simon D., Aden D.P., Knowles B.B.
Chromosomes of human hepatoma cell lines.
Int. J. Cancer 30:27-33(1982)
Patent=US4393133
Knowles B.B., Aden D.P.
Human hepatoma derived cell line, process for preparation thereof, and uses therefor.
Patent number US4393133, 12-Jul-1983
PubMed=2439335; DOI=10.1111/j.1432-1033.1987.tb11497.x
Vincent C., Marceau M., Blangarin P., Bouic P., Madjar J.J., Revillard J.-P.
Purification of alpha 1-microglobulin produced by human hepatoma cell lines. Biochemical characterization and comparison with alpha 1-microglobulin synthesized by human hepatocytes.
Eur. J. Biochem. 165:699-704(1987)
PubMed=8224613; DOI=10.1096/fasebj.7.14.8224613
Puisieux A., Galvin K., Troalen F., Bressac B., Marcais C., Galun E., Ponchel F., Yakicier C., Ji J.-W., Ozturk M.
Retinoblastoma and p53 tumor suppressor genes in human hepatoma cell lines.
FASEB J. 7:1407-1413(1993)
PubMed=8384076; DOI=10.1016/0165-4608(93)90227-D
Chen H.-L., Chiu T.-S., Chen P.-J., Chen D.-S.
Cytogenetic studies on human liver cancer cell lines.
Cancer Genet. Cytogenet. 65:161-166(1993)
PubMed=8389256; DOI=10.1093/carcin/14.5.987
Hsu I.C., Tokiwa T., Bennett W., Metcalf R.A., Welsh J.A., Sun T., Harris C.C.
p53 gene mutation and integrated hepatitis B viral DNA sequences in human liver cancer cell lines.
Carcinogenesis 14:987-992(1993)
PubMed=7972006; DOI=10.1073/pnas.91.23.11045
Okamoto A., Demetrick D.J., Spillare E.A., Hagiwara K., Hussain S.P., Bennett W.P., Forrester K., Gerwin B.I., Serrano M., Beach D.H., Harris C.C.
Mutations and altered expression of p16INK4 in human cancer.
Proc. Natl. Acad. Sci. U.S.A. 91:11045-11049(1994)
PubMed=8050184; DOI=10.1111/j.1365-2249.1994.tb06089.x
Wadee A.A., Paterson A., Coplan K.A., Reddy S.G.
HLA expression in hepatocellular carcinoma cell lines.
Clin. Exp. Immunol. 97:328-333(1994)
PubMed=8835345; DOI=10.1002/(SICI)1096-9071(199602)48:2<133::AID-JMV3>3.0.CO;2-A
Tsuboi S., Nagamori S., Miyazaki M., Mihara K., Fukaya K.-I., Teruya K., Kosaka T., Tsuji T., Namba M.
Persistence of hepatitis C virus RNA in established human hepatocellular carcinoma cell lines.
J. Med. Virol. 48:133-140(1996)
DOI=10.11418/jtca1981.16.3_173
Mihara K., Miyazaki M., Fushimi K., Tsuji T., Inoue Y., Fukaya K.-I., Ohashi R., Namba M.
The p53 gene status and other cellular characteristics of human cell lines maintained in our laboratory.
Tissue Cult. Res. Commun. 16:173-178(1997)
PubMed=9178645; DOI=10.1006/cimm.1997.1108
Nakao M., Sata M., Saitsu H., Yutani S., Kawamoto M., Kojiro M., Itoh K.
CD4+ hepatic cancer-specific cytotoxic T lymphocytes in patients with hepatocellular carcinoma.
Cell. Immunol. 177:176-181(1997)
PubMed=9359923; DOI=10.18926/AMO/30789
Mihara K., Miyazaki M., Kondo T., Fushimi K., Tsuji T., Inoue Y., Fukaya K.-I., Ishioka C., Namba M.
Yeast functional assay of the p53 gene status in human cell lines maintained in our laboratory.
Acta Med. Okayama 51:261-265(1997)
PubMed=11050057; DOI=10.1053/jhep.2000.19349
Wong N., Lai P., Pang E., Leung T.W.-T., Lau J.W.-L., Johnson P.J.
A comprehensive karyotypic study on human hepatocellular carcinoma by spectral karyotyping.
Hepatology 32:1060-1068(2000)
PubMed=11981770; DOI=10.1053/jhep.2002.32668
Clemens D.L., Forman A., Jerrells T.R., Sorrell M.F., Tuma D.J.
Relationship between acetaldehyde levels and cell survival in ethanol-metabolizing hepatoma cells.
Hepatology 35:1196-1204(2002)
PubMed=12029633; DOI=10.1053/jhep.2002.33683
Yasui K., Arii S., Zhao C., Imoto I., Ueda M., Nagai H., Emi M., Inazawa J.
TFDP1, CUL4A, and CDC16 identified as targets for amplification at 13q34 in hepatocellular carcinomas.
Hepatology 35:1476-1484(2002)
PubMed=12068308; DOI=10.1038/nature00766
Davies H., Bignell G.R., Cox C., Stephens P.J., Edkins S., Clegg S., Teague J.W., Woffendin H., Garnett M.J., Bottomley W., Davis N., Dicks E., Ewing R., Floyd Y., Gray K., Hall S., Hawes R., Hughes J., Kosmidou V., Menzies A., Mould C., Parker A., Stevens C., Watt S., Hooper S., Wilson R., Jayatilake H., Gusterson B.A., Cooper C.S., Shipley J.M., Hargrave D., Pritchard-Jones K., Maitland N.J., Chenevix-Trench G., Riggins G.J., Bigner D.D., Palmieri G., Cossu A., Flanagan A.M., Nicholson A., Ho J.W.C., Leung S.Y., Yuen S.T., Weber B.L., Seigler H.F., Darrow T.L., Paterson H.F., Marais R., Marshall C.J., Wooster R., Stratton M.R., Futreal P.A.
Mutations of the BRAF gene in human cancer.
Nature 417:949-954(2002)
DOI=10.1385/CP:1:3-4:313
Pang R.T.K., Poon T.C.-W., Wong N., Lai P.B.-S., Wong N.L.-Y., Chan C.M.-L., Yu J.W.S., Chan A.T.-C., Sung J.J.-Y.
Comparison of protein expression patterns between hepatocellular carcinoma cell lines and a hepatoblastoma cell line.
Clin. Proteomics 1:313-331(2004)
PubMed=14980111
Zhai B.-J., Wu F., Shao Z.-Y., Hu K., Zhao C.L., Wang Z.-B.
Establishment of human hepatocellular carcinoma multidrug-resistance cell line (HepG2/Adm) and study apoptosis induced by low-frequency pulse ultrasound exposure.
Zhonghua Gan Zang Bing Za Zhi 12:95-98(2004)
PubMed=15767549; DOI=10.1158/1535-7163.MCT-04-0234
Nakatsu N., Yoshida Y., Yamazaki K., Nakamura T., Dan S., Fukui Y., Yamori T.
Chemosensitivity profile of cancer cell lines and identification of genes determining chemosensitivity by an integrated bioinformatical approach using cDNA arrays.
Mol. Cancer Ther. 4:399-412(2005)
PubMed=16181800; DOI=10.1016/j.biocel.2005.07.010
Donohue T.M., Osna N.A., Clemens D.L.
Recombinant Hep G2 cells that express alcohol dehydrogenase and cytochrome P450 2E1 as a model of ethanol-elicited cytotoxicity.
Int. J. Biochem. Cell Biol. 38:92-101(2006)
PubMed=16935386; DOI=10.1016/j.jhep.2006.05.019
Sun D.-X., Nassal M.
Stable HepG2- and Huh7-based human hepatoma cell lines for efficient regulated expression of infectious hepatitis B virus.
J. Hepatol. 45:636-645(2006)
PubMed=17254797; DOI=10.1016/j.biologicals.2006.10.001
Azari S., Ahmadi N., Tehrani M.J., Shokri F.
Profiling and authentication of human cell lines using short tandem repeat (STR) loci: report from the National Cell Bank of Iran.
Biologicals 35:195-202(2007)
PubMed=19215227; DOI=10.1111/j.1349-7006.2009.01082.x
Kuwahara Y., Li L., Baba T., Nakagawa H., Shimura T., Yamamoto Y., Ohkubo Y., Fukumoto M.
Clinically relevant radioresistant cells efficiently repair DNA double-strand breaks induced by X-rays.
Cancer Sci. 100:747-752(2009)
PubMed=19751877; DOI=10.1016/j.humpath.2009.07.003
Lopez-Terrada D.H., Cheung S.W., Finegold M.J., Knowles B.B.
Hep G2 is a hepatoblastoma-derived cell line.
Hum. Pathol. 40:1512-1515(2009)
PubMed=20069059; DOI=10.1155/2010/437143
Srisomsap C., Sawangareetrakul P., Subhasitanont P., Chokchaichamnankit D., Chiablaem K., Bhudhisawasdi V., Wongkham S., Svasti J.
Proteomic studies of cholangiocarcinoma and hepatocellular carcinoma cell secretomes.
J. Biomed. Biotechnol. 2010:437143.1-437143.18(2010)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=20228232; DOI=10.1124/dmd.109.031831
Hart S.N., Li Y., Nakamoto K., Subileau E.-A., Steen D., Zhong X.-B.
A comparison of whole genome gene expression profiles of HepaRG cells and HepG2 cells to primary human hepatocytes and human liver tissues.
Drug Metab. Dispos. 38:988-994(2010)
PubMed=20937217; DOI=10.1170/149
Di Masi A., Viganotti M., Antoccia A., Magrelli A., Salvatore M., Azzalin G., Tosto F., Lorenzetti S., Maranghi F., Mantovani A., Macino G., Tanzarella C., Taruscio D.
Characterization of HuH6, Hep3B, HepG2 and HLE liver cancer cell lines by WNT/beta-catenin pathway, microRNA expression and protein expression profile.
Cell. Mol. Biol. 56:OL1299-OL1317(2010)
PubMed=21269460; DOI=10.1186/1752-0509-5-17
Burkard T.R., Planyavsky M., Kaupe I., Breitwieser F.P., Burckstummer T., Bennett K.L., Superti-Furga G., Colinge J.
Initial characterization of the human central proteome.
BMC Syst. Biol. 5:17.1-17.13(2011)
PubMed=22278370; DOI=10.1074/mcp.M111.014050
Geiger T., Wehner A., Schaab C., Cox J., Mann M.
Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins.
Mol. Cell. Proteomics 11:M111.014050-M111.014050(2012)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=23325432; DOI=10.1101/gr.147942.112
Varley K.E., Gertz J., Bowling K.M., Parker S.L., Reddy T.E., Pauli-Behn F., Cross M.K., Williams B.A., Stamatoyannopoulos J.A., Crawford G.E., Absher D.M., Wold B.J., Myers R.M.
Dynamic DNA methylation across diverse human cell lines and tissues.
Genome Res. 23:555-567(2013)
PubMed=23505090; DOI=10.1002/hep.26402
Wang K., Lim H.Y., Shi S., Lee J., Deng S., Xie T., Zhu Z., Wang Y., Pocalyko D., Yang W.J., Rejto P.A., Mao M., Park C.-K., Xu J.
Genomic landscape of copy number aberrations enables the identification of oncogenic drivers in hepatocellular carcinoma.
Hepatology 58:706-717(2013)
PubMed=23887712; DOI=10.1038/ncomms3218
Nault J.-C., Mallet M., Pilati C., Calderaro J., Bioulac-Sage P., Laurent C., Laurent A., Cherqui D., Balabaud C., Zucman-Rossi J.
High frequency of telomerase reverse-transcriptase promoter somatic mutations in hepatocellular carcinoma and preneoplastic lesions.
Nat. Commun. 4:2218.1-2218.7(2013)
PubMed=24116068; DOI=10.1371/journal.pone.0075692
Weiskirchen R., Weimer J., Meurer S.K., Kron A., Seipel B., Vater I., Arnold N., Siebert R., Xu L.-M., Friedman S.L., Bergmann C.
Genetic characteristics of the human hepatic stellate cell line LX-2.
PLoS ONE 8:E75692-E75692(2013)
PubMed=24618588; DOI=10.1371/journal.pone.0091433
Chernobrovkin A.L., Zubarev R.A.
Detection of viral proteins in human cells lines by xeno-proteomics: elimination of the last valid excuse for not testing every cellular proteome dataset for viral proteins.
PLoS ONE 9:E91433-E91433(2014)
PubMed=25960936; DOI=10.4161/21624011.2014.954893
Boegel S., Lower M., Bukur T., Sahin U., Castle J.C.
A catalog of HLA type, HLA expression, and neo-epitope candidates in human cancer cell lines.
OncoImmunology 3:e954893.1-e954893.12(2014)
PubMed=25485619; DOI=10.1038/nbt.3080
Klijn C., Durinck S., Stawiski E.W., Haverty P.M., Jiang Z.-S., Liu H.-B., Degenhardt J., Mayba O., Gnad F., Liu J.-F., Pau G., Reeder J., Cao Y., Mukhyala K., Selvaraj S.K., Yu M.-M., Zynda G.J., Brauer M.J., Wu T.D., Gentleman R.C., Manning G., Yauch R.L., Bourgon R., Stokoe D., Modrusan Z., Neve R.M., de Sauvage F.J., Settleman J., Seshagiri S., Zhang Z.-M.
A comprehensive transcriptional portrait of human cancer cell lines.
Nat. Biotechnol. 33:306-312(2015)
PubMed=25574106; DOI=10.3748/wjg.v21.i1.311
Cevik D., Yildiz G., Ozturk M.
Common telomerase reverse transcriptase promoter mutations in hepatocellular carcinomas from different geographical locations.
World J. Gastroenterol. 21:311-317(2015)
PubMed=26160117; DOI=10.1093/toxsci/kfv136
Sison-Young R.L.C., Mitsa D., Jenkins R.E., Mottram D., Alexandre E., Richert L., Aerts H., Weaver R.J., Jones R.P., Johann E., Hewitt P.G., Ingelman-Sundberg M., Goldring C.E.P., Kitteringham N.R., Park B.K.
Comparative proteomic characterization of 4 human liver-derived single cell culture models reveals significant variation in the capacity for drug disposition, bioactivation, and detoxication.
Toxicol. Sci. 147:412-424(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=26825538; DOI=10.1016/j.jprot.2016.01.016
Wisniewski J.R., Vildhede A., Noren A., Artursson P.
In-depth quantitative analysis and comparison of the human hepatocyte and hepatoma cell line HepG2 proteomes.
J. Proteomics 136:234-247(2016)
PubMed=27027780; DOI=10.1007/s10565-016-9316-2
Wu Y., Geng X.-C., Wang J.-F., Miao Y.-F., Lu Y.-L., Li B.
The HepaRG cell line, a superior in vitro model to L-02, HepG2 and hiHeps cell lines for assessing drug-induced liver injury.
Cell Biol. Toxicol. 32:37-59(2016)
PubMed=27329724; DOI=10.18632/oncotarget.10161
Watari K., Nishitani A., Shibata T., Noda M., Kawahara A., Akiba J., Murakami Y., Yano H., Kuwano M., Ono M.
Phosphorylation of mTOR Ser2481 is a key target limiting the efficacy of rapalogs for treating hepatocellular carcinoma.
Oncotarget 7:47403-47417(2016)
PubMed=27470094; DOI=10.1016/j.chroma.2016.07.042
Liu Z.-Y., Wang F.-J., Chen J., Zhou Y., Zou H.-F.
Modulating the selectivity of affinity absorbents to multi-phosphopeptides by a competitive substitution strategy.
J. Chromatogr. A 1461:35-41(2016)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=29610054; DOI=10.1016/j.dmpk.2018.03.003
Shi J., Wang X., Lyu L., Jiang H., Zhu H.-J.
Comparison of protein expression between human livers and the hepatic cell lines HepG2, Hep3B, and Huh7 using SWATH and MRM-HR proteomics: Focusing on drug-metabolizing enzymes.
Drug Metab. Pharmacokinet. 33:133-140(2018)
PubMed=29660373; DOI=10.1016/j.bbagen.2018.04.012
Touat-Hamici Z., Bulteau A.-L., Bianga J., Jean-Jacques H., Szpunar J., Lobinski R., Chavatte L.
Selenium-regulated hierarchy of human selenoproteome in cancerous and immortalized cells lines.
Biochim. Biophys. Acta 1862:2493-2505(2018)
PubMed=30629668; DOI=10.1371/journal.pone.0210404
Uphoff C.C., Pommerenke C., Denkmann S.A., Drexler H.G.
Screening human cell lines for viral infections applying RNA-Seq data analysis.
PLoS ONE 14:E0210404-E0210404(2019)
PubMed=30864654; DOI=10.1093/nar/gkz169
Zhou B., Ho S.S., Greer S.U., Spies N., Bell J.M., Zhang X., Zhu X., Arthur J.G., Byeon S., Pattni R., Saha I., Huang Y., Song G., Perrin D., Wong W.H., Ji H.P., Abyzov A., Urban A.E.
Haplotype-resolved and integrated genome analysis of the cancer cell line HepG2.
Nucleic Acids Res. 47:3846-3861(2019)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=31063779; DOI=10.1053/j.gastro.2019.05.001
Caruso S., Calatayud A.-L., Pilet J., La Bella T., Rekik S., Imbeaud S., Letouze E., Meunier L., Bayard Q., Rohr-Udilova N., Peneau C., Grasl-Kraupp B., de Koning L., Ouine B., Bioulac-Sage P., Couchy G., Calderaro J., Nault J.-C., Zucman-Rossi J., Rebouissou S.
Analysis of liver cancer cell lines identifies agents with likely efficacy against hepatocellular carcinoma and markers of response.
Gastroenterology 157:760-776(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. III, Barretina J.G., Gelfand E.T., Bielski C.M., Li H., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31378681; DOI=10.1016/j.ccell.2019.07.001
Qiu Z.-X., Li H., Zhang Z.-T., Zhu Z.-F., He S., Wang X.-J., Wang P.-C., Qin J.-J., Zhuang L.-P., Wang W., Xie F.-B., Gu Y., Zou K.-K., Li C., Li C., Wang C.-H., Cen J., Chen X.-T., Shu Y.-J., Zhang Z., Sun L.-L., Min L.-H., Fu Y., Huang X.-W., Lv H., Zhou H., Ji Y., Zhang Z.-G., Meng Z.-Q., Shi X.-L., Zhang H.-B., Li Y.-X., Hui L.-J.
A pharmacogenomic landscape in human liver cancers.
Cancer Cell 36:179-193.e11(2019)
PubMed=31395879; DOI=10.1038/s41467-019-11415-2
Yu K., Chen B., Aran D., Charalel J., Yau C., Wolf D.M., van 't Veer L.J., Butte A.J., Goldstein T., Sirota M.
Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types.
Nat. Commun. 10:3574.1-3574.11(2019)
PubMed=31561354; DOI=10.3233/CH-199226
Schulz C., Kammerer S., Kupper J.-H.
NADPH-cytochrome P450 reductase expression and enzymatic activity in primary-like human hepatocytes and HepG2 cells for in vitro biotransformation studies.
Clin. Hemorheol. Microcirc. 73:249-260(2019)
PubMed=31978347; DOI=10.1016/j.cell.2019.12.023
Nusinow D.P., Szpyt J., Ghandi M., Rose C.M., McDonald E.R. III, Kalocsay M., Jane-Valbuena J., Gelfand E.T., Schweppe D.K., Jedrychowski M.P., Golji J., Porter D.A., Rejtar T., Wang Y.K., Kryukov G.V., Stegmeier F., Erickson B.K., Garraway L.A., Sellers W.R., Gygi S.P.
Quantitative proteomics of the Cancer Cell Line Encyclopedia.
Cell 180:387-402.e16(2020)
PubMed=32471280; DOI=10.3390/ijms21113813
Kiseleva O., Ponomarenko E., Poverennaya E.
Empowering shotgun mass spectrometry with 2DE: a HepG2 study.
Int. J. Mol. Sci. 21:3813.1-3813.16(2020)
PubMed=32899426; DOI=10.3390/cancers12092510
Scherer D., Davila Lopez M., Goeppert B., Abrahamsson S., Gonzalez Silos R., Nova I., Marcelain K., Roa J.C., Ibberson D., Umu S.U., Rounge T.B., Roessler S., Bermejo J.L.
RNA sequencing of hepatobiliary cancer cell lines: data and applications to mutational and transcriptomic profiling.
Cancers (Basel) 12:2510.1-2510.14(2020)
PubMed=34320349; DOI=10.1016/j.celrep.2021.109441
Jochems F., Thijssen B., De Conti G., Jansen R., Pogacar Z., Groot K., Wang L.-Q., Schepers A., Wang C., Jin H.-J., Beijersbergen R.L., Leite de Oliveira R., Wessels L.F.A., Bernards R.
The cancer SENESCopedia: a delineation of cancer cell senescence.
Cell Rep. 36:109441.1-109441.22(2021)
 技术资料
技术资料
需要更多技术资料 索取更多技术资料


