pET28GST-LIC
PVT10579 2ug
pET28GST-LIC Information
Promoter: T7/Laco promoter
Replicon: pBR322 ori, F1 ori
Terminator: T7 Terminator
Plasmid size: 7994bp
Plasmid tagging: N-GST, N-6 x HIS, N-thrombin site, SacB
Prokaryotic resistance: kanamycin Kan
Screening marker: SacB/ sucrose screening
Clone strain: Escherichia coli DH5 alpha
Culture conditions: 37 C, aerobic, LB
Expression vector: Escherichia coli BL21 (DE3)
Culture conditions: 37 C, aerobic, LB
Induction: IPTG and lactose and its analogues.
5'sequencing primers: T7 (TAATACGACTCACTATAGGG)
3'sequencing primers: T7-ter (TGCTAGTTATTGCTCAGCGG)
pET28GST-LIC Description
pET28GST-LIC is an E.coli expression vector. T7 promoter drives GST to express with fusion expression. SacB lethal gene is on the vector and positive clones can be screened by 5% sucrose.
pET28GST-LIC vector was derived from expression plasmid pET28a-LIC (SGC) by inserting the GST-tag from pET41a (Novagen) into the XbaI and NcoI sites. It is used for T7 promoter driven expression of recombinant proteins with the addition of a 242 amino acid N-terminal fusion tag containing the 217 amino acid GST-tag protein followed by a 6X His followed by a thrombin cleavage site. Two stop codons are included in the vector at the C-terminal cloning site.Insertion of DNA sequence into the cloning/expression region is preformed using BD-Biosciences Infusion enzyme mediated directional recombination between complementary 15 nucleotide DNA sequences at the ends of the insert (PCR product) and BseRI linearized vector. Insertion of target sequence involves replacement of a SacB gene stuffer sequence, which provides for negative selection of the original plasmid on 5% sucrose.
pET28GST-LIC Multiple cloning site
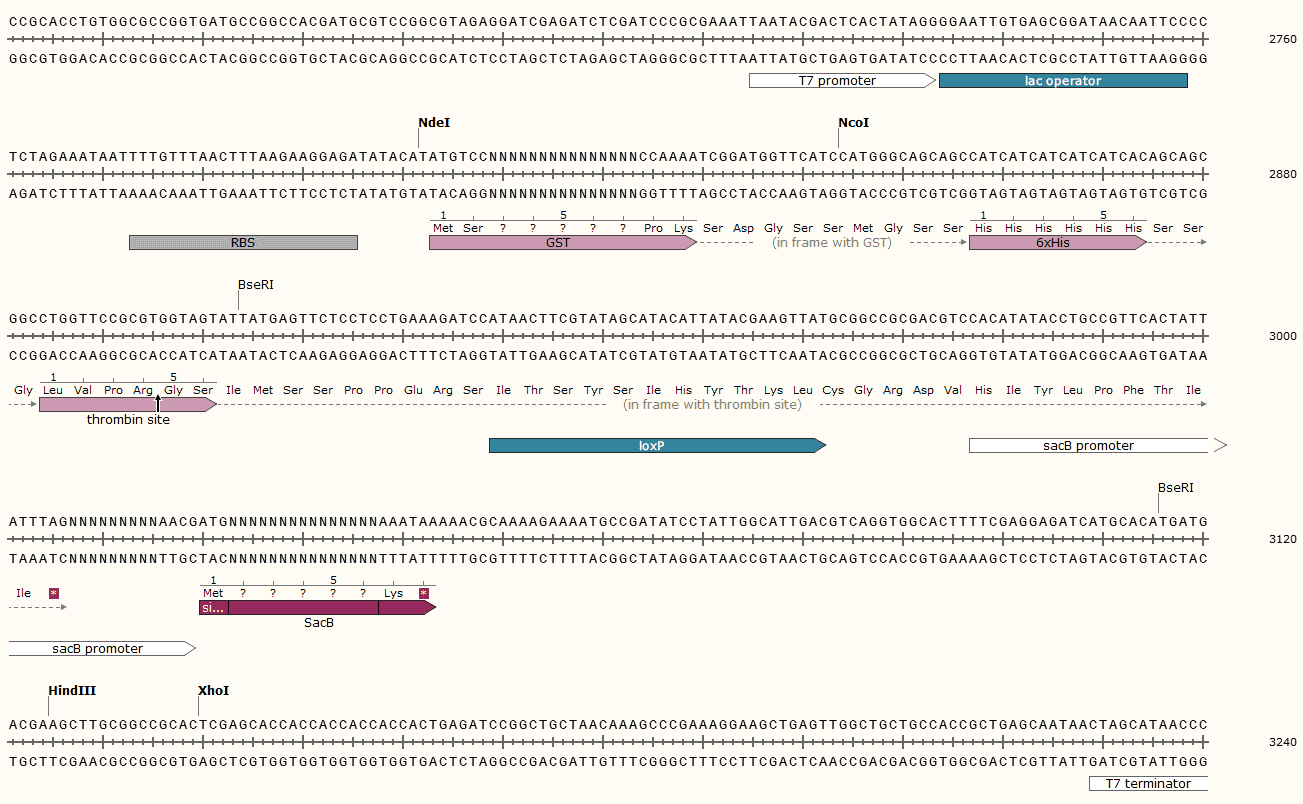
pET28GST-LIC Sequence
LOCUS Exported 7998 bp ds-DNA circular SYN 17-NOV-2017
DEFINITION synthetic circular DNA
SOURCE synthetic DNA construct
ORGANISM synthetic DNA construct
REFERENCE 1 (bases 1 to 7998)
AUTHORS .
TITLE Direct Submission
FEATURES Location/Qualifiers
source 1..7998
/organism="synthetic DNA construct"
/mol_type="other DNA"
rep_origin 12..467
/direction=RIGHT
/label=f1 ori
/note="f1 bacteriophage origin of replication; arrow
indicates direction of (+) strand synthesis"
CDS complement(560..1375)
/codon_start=1
/gene="aph(3')-Ia"
/product="aminoglycoside phosphotransferase"
/label=KanR
/note="confers resistance to kanamycin in bacteria or G418
(Geneticin(R)) in eukaryotes"
/translation="MSHIQRETSCSRPRLNSNMDADLYGYKWARDNVGQSGATIYRLYG
KPDAPELFLKHGKGSVANDVTDEMVRLNWLTEFMPLPTIKHFIRTPDDAWLLTTAIPGK
TAFQVLEEYPDSGENIVDALAVFLRRLHSIPVCNCPFNSDRVFRLAQAQSRMNNGLVDA
SDFDDERNGWPVEQVWKEMHKLLPFSPDSVVTHGDFSLDNLIFDEGKLIGCIDVGRVGI
ADRYQDLAILWNCLGEFSPSLQKRLFQKYGIDNPDMNKLQFHLMLDEFF"
rep_origin 1497..2085
/direction=RIGHT
/label=ori
/note="high-copy-number ColE1/pMB1/pBR322/pUC origin of
replication"
misc_feature 2271..2413
/label=bom
/note="basis of mobility region from pBR322"
CDS complement(2515..2706)
/codon_start=1
/gene="rop"
/product="Rop protein, which maintains plasmids at low copy
number"
/label=rop
/translation="MTKQEKTALNMARFIRSQTLTLLEKLNELDADEQADICESLHDHA
DELYRSCLARFGDDGENL"
CDS complement(3515..4597)
/codon_start=1
/gene="lacI"
/product="lac repressor"
/label=lacI
/note="The lac repressor binds to the lac operator to
inhibit transcription in E. coli. This inhibition can be
relieved by adding lactose or
isopropyl-beta-D-thiogalactopyranoside (IPTG)."
/translation="MKPVTLYDVAEYAGVSYQTVSRVVNQASHVSAKTREKVEAAMAEL
NYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKSRADQLGASVVVSMVERSGV
EACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNVPALFLDVSDQTPINSIIFSH
EDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGWHKYLTRNQIQPIAEREGDWSA
MSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITESGLRVGADISVVGYDDTEDSSC
YIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQLLPVSLVKRKTTLAPNTQTASPR
ALADSLMQLARQVSRLESGQ"
promoter complement(4598..4675)
/gene="lacI"
/label=lacI promoter
promoter 4984..5002
/label=T7 promoter
/note="promoter for bacteriophage T7 RNA polymerase"
protein_bind 5003..5027
/label=lac operator
/bound_moiety="lac repressor encoded by lacI"
/note="The lac repressor binds to the lac operator to
inhibit transcription in E. coli. This inhibition can be
relieved by adding lactose or
isopropyl-beta-D-thiogalactopyranoside (IPTG)."
RBS 5042..5064
/note="efficient ribosome binding site from bacteriophage
T7 gene 10 (Olins and Rangwala, 1989)"
CDS 5072..5725
/codon_start=1
/product="glutathione S-transferase from Schistosoma
japonicum"
/label=GST
/translation="MSPILGYWKIKGLVQPTRLLLEYLEEKYEEHLYERDEGDKWRNKK
FELGLEFPNLPYYIDGDVKLTQSMAIIRYIADKHNMLGGCPKERAEISMLEGAVLDIRY
GVSRIAYSKDFETLKVDFLSKLPEMLKMFEDRLCHKTYLNGDHVTHPDFMLYDALDVVL
YMDPMCLDAFPKLVCFKKRIEAIPQIDKYLKSSKYIAWPLQGWQATFGGGDHPPK"
CDS 5753..5770
/codon_start=1
/product="6xHis affinity tag"
/label=6xHis
/translation="HHHHHH"
CDS 5780..5797
/codon_start=1
/product="thrombin recognition and cleavage site"
/label=thrombin site
/translation="LVPRGS"
protein_bind 5825..5858
/label=loxP
/bound_moiety="Cre recombinase"


 文献和实验
文献和实验


