pER8
Search name
pER8,Plasmid pER8,pER8 vector
pER8 Plasmid Information
Promoter: CaMV 35S
Replicon: ori
Terminator: NOS
Plasmid classification: plant series, protein overexpression vector
Plasmid size: 11522bp
Prokaryotic resistance: Spe
Selection marker: HygR
Clone strain: DH5 alpha
Culture conditions: 37 LB, aerobic
Expressing host: plant cell
Primers for 5'sequencing: M13F:TGTAAAACGACGGCCAGT
Primers for 3'sequencing: primers were designed according to the sequence
Use: Plant expression
pER8 Plasmid Sequence
LOCUS Exported 11522 bp ds-DNA circular SYN 11-JUL-2015
DEFINITION synthetic circular DNA
ACCESSION .
VERSION .
KEYWORDS pER8
SOURCE synthetic DNA construct
ORGANISM synthetic DNA construct
REFERENCE 1 (bases 1 to 11522)
AUTHORS Gang Yu
TITLE Direct Submission
JOURNAL Exported Wednesday, September 14, 2016 from SnapGene Viewer 3.2.1
FEATURES Location/Qualifiers
source 1..11522
/organism="synthetic DNA construct"
/mol_type="other DNA"
misc_feature 1..25
/note="RB T-DNA repeat"
/note="right border repeat from nopaline C58 T-DNA"
promoter 216..262
/note="minimal CaMV 35S promoter"
/note="minimal 35S promoter from cauliflower mosaic virus"
CDS 291..542
/codon_start=1
/gene="lexA (truncated)"
/product="DNA-binding domain of E. coli LexA"
/note="LexA BD"
/note="binds the 16-bp palindromic sequence
5'-CTGTATATATATACAG-3'"
/translation="MKALTARQQEVFDLIRDHISQTGMPPTRAEIAQRLGFRSPNAAEE
HLKALARKGVIEIVSGASRGIRLLQEEEEGLPLVGRVAA"
CDS 558..791
/codon_start=1
/gene="UL48"
/product="transcriptional activation domain of herpes
simplex virus protein VP16 (Triezenberg et al., 1988;
Cousens et al., 1989)"
/note="VP16 AD"
/translation="APPTDVSLGDELHLDGEDVAMAHADALDDFDLDMLGDGDSPGPGF
TPHDSAPYGALDMADFEFEQMFTDALGIDEYGG"
CDS 849..1742
/codon_start=1
/gene="ESR1"
/product="estrogen nuclear receptor alpha ligand-binding
domain"
/note="ER-LBD"
/translation="KRSKKNSLALSLTADQMVSALLDAEPPILYSEYDPTRPFSEASMM
GLLTNLADRELVHMINWAKRVPGFVDLTLHDQVHLLECAWLEILMIGLVWRSMEHPVKL
LFAPNLLLDRNQGKCVEGMVEIFDMLLATSSRFRMMNLQGEEFVCLKSIILLNSGVYTF
LSSTLKSLEEKDHIHRVLDKITDTLIHLMAKAGLTLQQQHQRLAQLLLILSHIRHMSNK
GMEHLYSMKCKNVVPLYDLLLEMLDAHRLHAPTSRGGASVEETDQSHLATAGSTSSHSL
QKYYITGEAEGFPATV"
promoter 2294..2474
/note="NOS promoter"
/note="nopaline synthase promoter"
promoter 2327..2510
/function="NOS promoter"
/note="NOS promoter"
/note="nopaline synthase promoter"
CDS 2525..3550
/codon_start=1
/gene="aph(4)-Ia"
/product="aminoglycoside phosphotransferase from E. coli"
/note="HygR"
/note="confers resistance to hygromycin"
/translation="MKKPELTATSVEKFLIEKFDSVSDLMQLSEGEESRAFSFDVGGRG
YVLRVNSCADGFYKDRYVYRHFASAALPIPEVLDIGEFSESLTYCISRRAQGVTLQDLP
ETELPAVLQPVAEAMDAIAAADLSQTSGFGPFGPQGIGQYTTWRDFICAIADPHVYHWQ
TVMDDTVSASVAQALDELMLWAEDCPEVRHLVHADFGSNNVLTDNGRITAVIDWSEAMF
GDSQYEVANIFFWRPWLACMEQQTRYFERRHPELAGSPRLRAYMLRIGLDQLYQSLVDG
NFDDAAWAQGRCDAIVRSGAGTVGRTQIARRSAAVWTDGCVEVLADSGNRRPSTRPRAK
E"
terminator 3552..3804
/note="NOS terminator"
/note="nopaline synthase terminator and poly(A) signal"
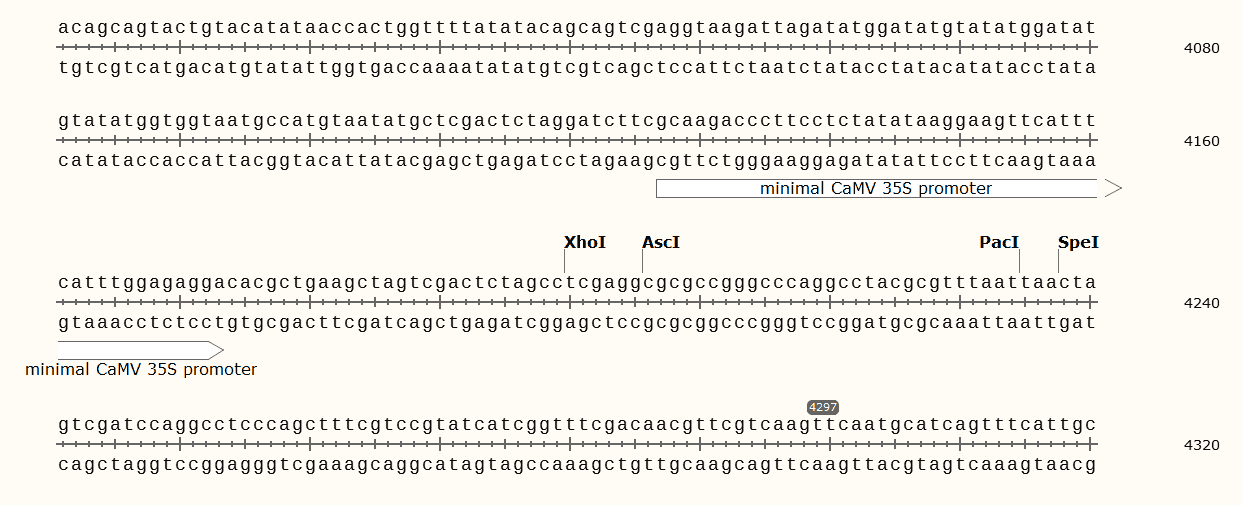
promoter 4127..4173
/note="minimal CaMV 35S promoter"
/note="minimal 35S promoter from cauliflower mosaic virus"
primer_bind complement(4750..4766)
/note="M13 fwd"
/note="common sequencing primer, one of multiple similar
variants"
misc_feature 4988..5012
/note="LB T-DNA repeat"
/note="left border repeat from nopaline C58 T-DNA"
CDS 5540..6331
/codon_start=1
/gene="aadA"
/product="aminoglycoside adenylyltransferase (Murphy,
1985)"
/note="SmR"
/note="confers resistance to spectinomycin and
streptomycin"
/translation="MGEAVIAEVSTQLSEVVGVIERHLEPTLLAVHLYGSAVDGGLKPH
SDIDLLVTVTVRLDETTRRALINDLLETSASPGESEILRAVEVTIVVHDDIIPWRYPAK
RELQFGEWQRNDILAGIFEPATIDIDLAILLTKAREHSVALVGPAAEELFDPVPEQDLF
EALNETLTLWNSPPDWAGDERNVVLTLSRIWYSAVTGKIAPKDVAADWAMERLPAQYQP
VILEARQAYLGQEEDRLASRADQLEEFVHYVKGEITKVVGK"
rep_origin 6575..7163
/direction=RIGHT
/note="ori"
/note="high-copy-number ColE1/pMB1/pBR322/pUC origin of


 文献和实验
文献和实验


