pCAMBIA2300
Search name
pCAMBIA2300,Plasmid pCAMBIA2300,pCAMBIA2300 vector
pCAMBIA2300 Informaiton
Promoter: Lac, CaMV 35S
Replicon: pVS1 oriV, ori
Plasmid classification: plant series, protein overexpression vector
Plasmid size: 8743bp
Prokaryotic resistance: Kan
Selection marker: Neo
Clonal strain: DH5 alpha
Culture conditions: 37 C, aerobic LB
Expression host: plant cells
5'sequencing primers: 35S:GACGCACAATCCCACTATCC
Primers for 3'sequencing: primers designed based on sequences
Use: Plant expression
pCAMBIA2300 Description
The pCambia2300 vector offers: 1.High copy number in E.coli for high DNA yields 2. pVS1 replicon for high stability in Agrobacterium3.Small size 4.Restriction sites designed for modular plasmid modifications and small but adequate poly-linkers for introducing your DNA of interest 5. Bacterial selection with kanamycin
pCAMBIA2300 Multiple cloning site
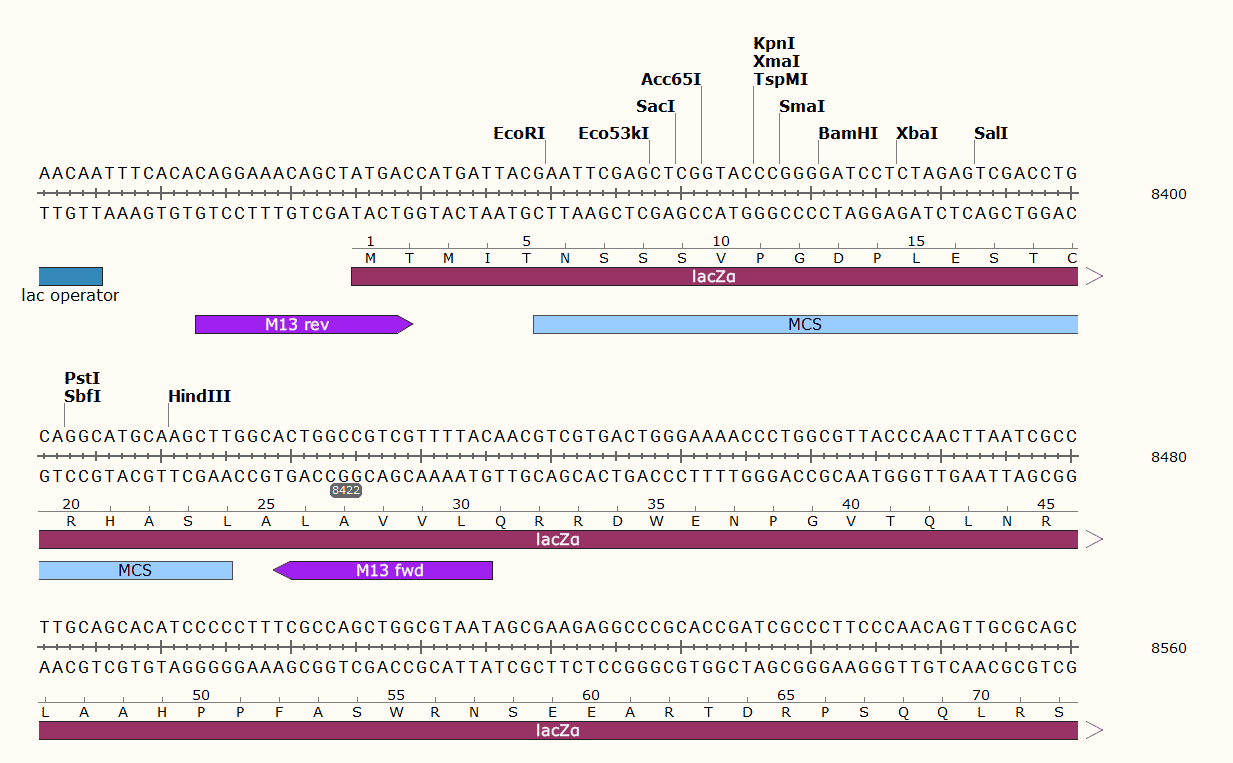
pCAMBIA2300 Sequence
LOCUS Exported 8743 bp ds-DNA circular SYN 11-JAN-2016
DEFINITION Agrobacterium binary vector for plant transformation, with
kanamycin-resistance genes.
ACCESSION AF234315
VERSION .
KEYWORDS pCAMBIA2300
SOURCE synthetic DNA construct
ORGANISM synthetic DNA construct
REFERENCE 1 (bases 1 to 8743)
AUTHORS Cambia
TITLE Direct Submission
JOURNAL Exported Wednesday, September 14, 2016 from SnapGene Viewer 3.2.1
COMMENT The GenBank record was corrected by inserting a G at position 2281.
FEATURES Location/Qualifiers
source 1..8743
/organism="synthetic DNA construct"
/lab_host="Plant Cells"
/mol_type="other DNA"
CDS 1219..1848
/codon_start=1
/product="stability protein from Pseudomonas plasmid pVS1"
/note="pVS1 StaA"
/translation="MKVIAVLNQKGGSGKTTIATHLARALQLAGADVLLVDSDPQGSAR
DWAAVREDQPLTVVGIDRPTIDRDVKAIGRRDFVVIDGAPQAADLAVSAIKAADFVLIP
VQPSPYDIWATADLVELVKQRIEVTDGRLQAAFVVSRAIKGTRIGGEVAEALAGYELPI
LESRITQRVSYPGTAAAGTTVLESEPEGDAAREVQALAAEIKSKLI"
CDS 2277..3350
/codon_start=1
/product="replication protein from Pseudomonas plasmid
pVS1"
/note="pVS1 RepA"
/translation="MSGRKPSGPVQIGAALGDDLVEKLKAAQAAQRQRIEAEARPGESW
QAAADRIRKESRQPPAAGAPSIRKPPKGDEQPDFFVPMLYDVGTRDSRSIMDVAVFRLS
KRDRRAGEVIRYELPDGHVEVSAGPAGMASVWDYDLVLMAVSHLTESMNRYREGKGDKP
GRVFRPHVADVLKFCRRADGGKQKDDLVETCIRLNTTHVAMQRTKKAKNGRLVTVSEGE
ALISRYKIVKSETGRPEYIEIELADWMYREITEGKNPDVLTVHPDYFLIDPGIGRFLYR
LARRAAGKAEARWLFKTIYERSGSAGEFKKFCFTVRKLIGSNDLPEYDLKEEAGQAGPI
LVMRYRNLIEGEASAGS"
rep_origin 3416..3610
/note="pVS1 oriV"
/note="origin of replication for the Pseudomonas plasmid
pVS1 (Heeb et al., 2000)"
misc_feature 3954..4094
/note="bom"
/note="basis of mobility region from pBR322"
rep_origin complement(4280..4868)
/direction=LEFT
/note="ori"
/note="high-copy-number ColE1/pMB1/pBR322/pUC origin of
replication"
CDS complement(4955..5749)
/codon_start=1
/gene="aphA-3"
/product="aminoglycoside phosphotransferase"
/note="KanR"
/note="confers resistance to kanamycin"
/translation="MAKMRISPELKKLIEKYRCVKDTEGMSPAKVYKLVGENENLYLKM
TDSRYKGTTYDVEREKDMMLWLEGKLPVPKVLHFERHDGWSNLLMSEADGVLCSEEYED
EQSPEKIIELYAECIRLFHSIDISDCPYTNSLDSRLAELDYLLNNDLADVDCENWEEDT
PFKDPRELYDFLKTEKPEEELVFSHGDLGDSNIFVKDGKVSGFIDLGRSGRADKWYDIA
FCVRSIREDIGEEQYVELFFDLLGIKPDWEKIKYYILLDELF"
misc_feature 6174..6198
/note="LB T-DNA repeat"
/note="left border repeat from nopaline C58 T-DNA"
polyA_signal 6276..6450
/note="CaMV poly(A) signal"
/note="cauliflower mosaic virus polyadenylation signal"
CDS complement(6507..7304)
/codon_start=1
/gene="aph(3')-II (or nptII)"
/product="aminoglycoside phosphotransferase from Tn5"
/note="NeoR/KanR"
/note="confers resistance to neomycin, kanamycin, and G418
(Geneticin(R))"
/translation="MGIEQDGLHAGSPAAWVERLFGYDWAQQTIGCSDAAVFRLSAQGR
PVLFVKTDLSGALNELQDEAARLSWLATTGVPCAAVLDVVTEAGRDWLLLGEVPGQDLL
SSHLAPAEKVSIMADAMRRLHTLDPATCPFDHQAKHRIERARTRMEAGLVDQDDLDEEH
QGLAPAELFARLKARMPDGEDLVVTHGDACLPNIMVENGRFSGFIDCGRLGVADRYQDI
ALATRDIAEELGGEWADRFLVLYGIAAPDSQRIAFYRLLDEFF"
promoter complement(7367..8044)
/note="CaMV 35S promoter (enhanced)"
/note="cauliflower mosaic virus 35S promoter with a
duplicated enhancer region"
promoter 8271..8301
/note="lac promoter"
/note="promoter for the E. coli lac operon"
protein_bind 8309..8325
/bound_moiety="lac repressor encoded by lacI"
/note="lac operator"
/note="The lac repressor binds to the lac operator to
inhibit transcription in E. coli. This inhibition can be
relieved by adding lactose or
isopropyl-beta-D-thiogalactopyranoside (IPTG)."
primer_bind 8333..8349
/note="M13 rev"
/note="common sequencing primer, one of multiple similar
variants"
CDS 8345..8578
/codon_start=1
/gene="lacZ (truncated)"


 文献和实验
文献和实验


