pLVX-AcGFP1-N1
Search name
pLVX-AcGFP1-N1,Plasmid pLVX-AcGFP1-N1,pLVX-AcGFP1-N1 vector
pLVX-AcGFP1-N1 Information
Promoter: PGK, CMV
Replicon: pUC ori
Terminator: SV40 poly (A) signal
Plasmid Classification: Virus Series, Lentivirus Cloning Vectors
Plasmid size: 8787bp
Prokaryotic resistance: Amp
Selection marker: Puro
Clone strain: Stbl3
Culture conditions: 37 C, aerobic LB
Expression host: mammalian cells
Induction mode: no induction, transient expression
5'Sequencing Primers: CMV-F: CGCAAATGGGGGGGTAGGGGGGTG
3'Sequencing Primers: Designing Primers Based on Sequences
pLVX-AcGFP1-N1 Description
Plvx-acgfp1-n1 is HIV-1, a lentivirus expression vector.
The gene was cloned into the polyclonal site (MCS), and the coding sequence of ACGFP1 was located upstream. The expression of the N-terminal fusion protein of ACGFP1 was expressed by the human cytomegalovirus-driven early promoter (CMV) located upstream. Lentivirus particles come from vectors that allow AcGFP1 fusion protein to be expressed in almost any type of cell, including primary cells. The unmodifi ED vector represents AcGFP1 and can be used to optimize the infection protocol of production marker virus.
pLVX-AcGFP1-N1 Multiple cloning sites
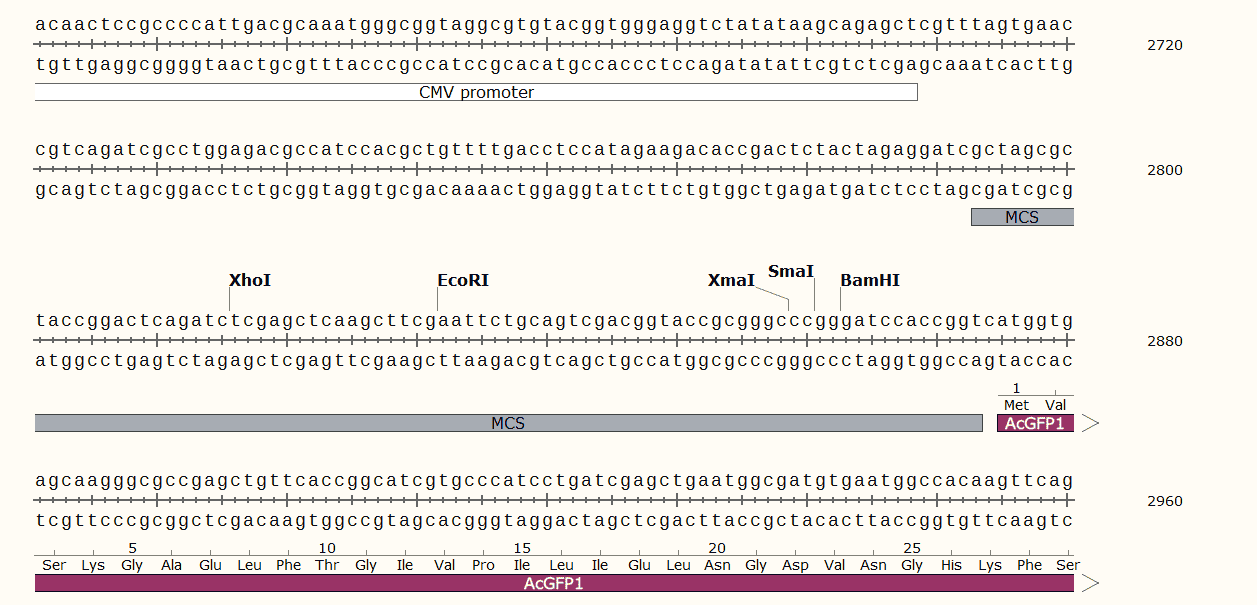

pLVX-AcGFP1-N1 Sequence
LOCUS Exported 8787 bp ds-DNA circular SYN 27-AUG-2016
DEFINITION synthetic circular DNA
ACCESSION .
VERSION .
KEYWORDS Untitled 23
SOURCE synthetic DNA construct
ORGANISM synthetic DNA construct
REFERENCE 1 (bases 1 to 8787)
AUTHORS .
TITLE Direct Submission
JOURNAL Exported Saturday, August 27, 2016 from SnapGene Viewer 3.1.4
FEATURES Location/Qualifiers
source 1..8787
/organism="synthetic DNA construct"
/mol_type="other DNA"
LTR 1..634
/note="3' LTR"
/note="3' long terminal repeat (LTR) from HIV-1"
misc_feature 681..806
/note="HIV-1 Psi"
/note="packaging signal of human immunodeficiency virus
type 1"
misc_feature 1303..1536
/note="RRE"
/note="The Rev response element (RRE) of HIV-1 allows for
Rev-dependent mRNA export from the nucleus to the
cytoplasm."
misc_feature 2028..2143
/note="cPPT/CTS"
/note="central polypurine tract and central termination
sequence of HIV-1 (lacking the first T)"
enhancer 2201..2504
/note="CMV enhancer"
/note="human cytomegalovirus immediate early enhancer"
promoter 2505..2708
/note="CMV promoter"
/note="human cytomegalovirus (CMV) immediate early
promoter"
misc_feature 2793..2873
/note="MCS"
/note="multiple cloning site"
CDS 2875..3594
/codon_start=1
/product="Aequorea coerulescens GFP"
/note="AcGFP1"
/note="mammalian codon-optimized"
/translation="MVSKGAELFTGIVPILIELNGDVNGHKFSVSGEGEGDATYGKLTL
KFICTTGKLPVPWPTLVTTLSYGVQCFSRYPDHMKQHDFFKSAMPEGYIQERTIFFEDD
GNYKSRAEVKFEGDTLVNRIELTGTDFKEDGNILGNKMEYNYNAHNVYIMTDKAKNGIK
VNFKIRHNIEDGSVQLADHYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEKRDHMIYF
GFVTAAAITHGMDELYK"
promoter 3622..4121
/note="PGK promoter"
/note="mouse phosphoglycerate kinase 1 promoter"
CDS 4142..4741
/codon_start=1
/gene="pac from Streptomyces alboniger"
/product="puromycin N-acetyltransferase"
/note="PuroR"
/note="confers resistance to puromycin"
/translation="MTEYKPTVRLATRDDVPRAVRTLAAAFADYPATRHTVDPDRHIER
VTELQELFLTRVGLDIGKVWVADDGAAVAVWTTPESVEAGAVFAEIGPRMAELSGSRLA
AQQQMEGLLAPHRPKEPAWFLATVGVSPDHQGKGLGSAVVLPGVEAAERAGVPAFLETS
APRNLPFYERLGFTVTADVEVPEGPRTWCMTRKPGA"
misc_feature 4755..5343
/note="WPRE"
/note="woodchuck hepatitis virus posttranscriptional
regulatory element"
CDS complement(5226..5237)
/codon_start=1
/product="Factor Xa recognition and cleavage site"
/note="Factor Xa site"
/translation="IEGR"
LTR 5550..6183
/note="3' LTR"
/note="3' long terminal repeat (LTR) from HIV-1"
primer_bind complement(6312..6328)
/note="M13 rev"
/note="common sequencing primer, one of multiple similar
variants"
protein_bind 6336..6352
/bound_moiety="lac repressor encoded by lacI"
/note="lac operator"
/note="The lac repressor binds to the lac operator to
inhibit transcription in E. coli. This inhibition can be
relieved by adding lactose or
isopropyl-beta-D-thiogalactopyranoside (IPTG)."
promoter complement(6360..6390)
/note="lac promoter"
/note="promoter for the E. coli lac operon"
protein_bind 6405..6426
/bound_moiety="E. coli catabolite activator protein"
/note="CAP binding site"
/note="CAP binding activates transcription in the presence
of cAMP."
rep_origin complement(6714..7302)
/direction=LEFT
/note="ori"
/note="high-copy-number ColE1/pMB1/pBR322/pUC origin of
replication"
CDS complement(7473..8333)


 文献和实验
文献和实验