万千商家帮你免费找货
0 人在求购买到急需产品
- 详细信息
- 用户评价
- 文献和实验
- 技术资料
- 服务名称:
多肽合成,多肽定制
- 提供商:
北京中科亚光生物科技有限公司
- 规格:
1mg,5mg,10mg等
- 标准肽:链长至 100个氨基酸,毫克至克级,纯度可高达99%。
- 不同纯度范围:粗肽、脱盐、>75% 、> 85% 、>90%、>95%、>98%、>99%。
- 常规修饰肽:乙酰化,酰胺化,生物素标记肽,磷酸化肽,D型氨基酸修饰肽等。
- 特殊修饰肽:磺酸化肽(Sulfated-tyrosine),环肽(二硫键环化,首尾环化),荧光标记肽(FITC,Dabcyl/Edans,Dansyl,FAM,Abz/Dnp,Rhodamine等),甲基化多肽(Lys(Me2),Lys(Me),Lys(Me3),Arg(Me),Arg(Me2) -Symetrical,Arg(Me2) –Asymetrical),同位素标记肽(Heavy Isotope Labeled Peptides)、拟肽(peptoid)和点击化学用多肽(Peptides for Click Chemistry)。
- 药用肽,大批量肽。
- 协助客户建立科研肽库。
- 抗原多肽及其与蛋白的交联:KLH,BSA, OVA, HEL。
服务流程
- 确定合成序列,我们将在12小时内容提供准确报价。
- 签订委托合同及保密协议。
- 根据合同内容,在规定时间内交货,货到付款。
- 对于合成的每条多肽,将免费提供HPLC和MS检测报告。
- 产品以冻干粉状态包装,以快递方式送达客户手中(顺丰包邮)。
服务说明
- 基酸残基数少于6个的多肽以6个氨基酸计。氨基酸残基数大于30个的多肽欢迎来电咨询。
- 我们提供的产品质量范围在毫克级至百克级之间,以充分满足客户的不同需求。
- 100条或100条以下的普通多肽粗品会在1-2周内交货,纯化产品将在2-3周内交货;100条以上或者超过30条氨基酸残基的复杂多肽交货时间相应延长。
- 对于每一条多肽,我们在向客户提供优质产品的同时,提供高标准的质检报告(包括高效液相色谱分析图,质谱图)。
- 我们将为客户合成的多肽序列严格保密。在确认多肽序列后,我们可以与客户签订保密协议。
- 在充分考虑到客户研发预算的前提下,我们将以优惠的价格最大限度的满足客户的实验需求,欢迎随时与我们联系。

亚光多肽合成服务产品优势
| 质量关做到每个过程有记录可查,严格质量标准,为客户实验的可靠性提供保障。 |
| 多肽的高通量合成极大降低综合成本,为您赢得更多时间,降低科研费用。 |
| 在修饰肽、标记肽以及特殊结构多肽等多种复杂多肽合成技术上精益求精。 |
| 专业的售前和售后服务,让用户在多肽咨询、订购以及使用方面无后顾之忧。 |
使用我公司多肽产品发表的代表性文章:
[1] Molecular basis for histone N-terminal methylation by NRMT1, Genes & Development, 2015, 29: 2337-2342.
[2] ZMYND11 links histone H3.3K36me3 to transcription elongation and tumour suppression, Nature, 2014, 508: 263-268.
[3] The MTA family proteins as novel histone H3 binding proteins, Cell Biosci, 2013, 3(1): 1.
[4] Short-Form CDYLb but not long-form CDYLa functions cooperatively with histone methyltransferase G9a in hepatocellular carcinomas, Genes Chromosomes Cancer, 2013, 52(7): 644-655.
[5] Global profiling of regulatory elements in the histone benzoylation pathway, NATURE COMMUNICATIONS, 2022,https://doi.org/10.1038/s41467-022-29057-2.
公司实力
引进国际先进的多肽合成技术,并且配备先进全自动多通道合成仪,同时合成上百条多肽,每月生产能力达上千条。在现有的多肽合成技术基础上,我们对其进行了优化和改造,已经建立较为完善的高通量大规模多肽技术服务平台。
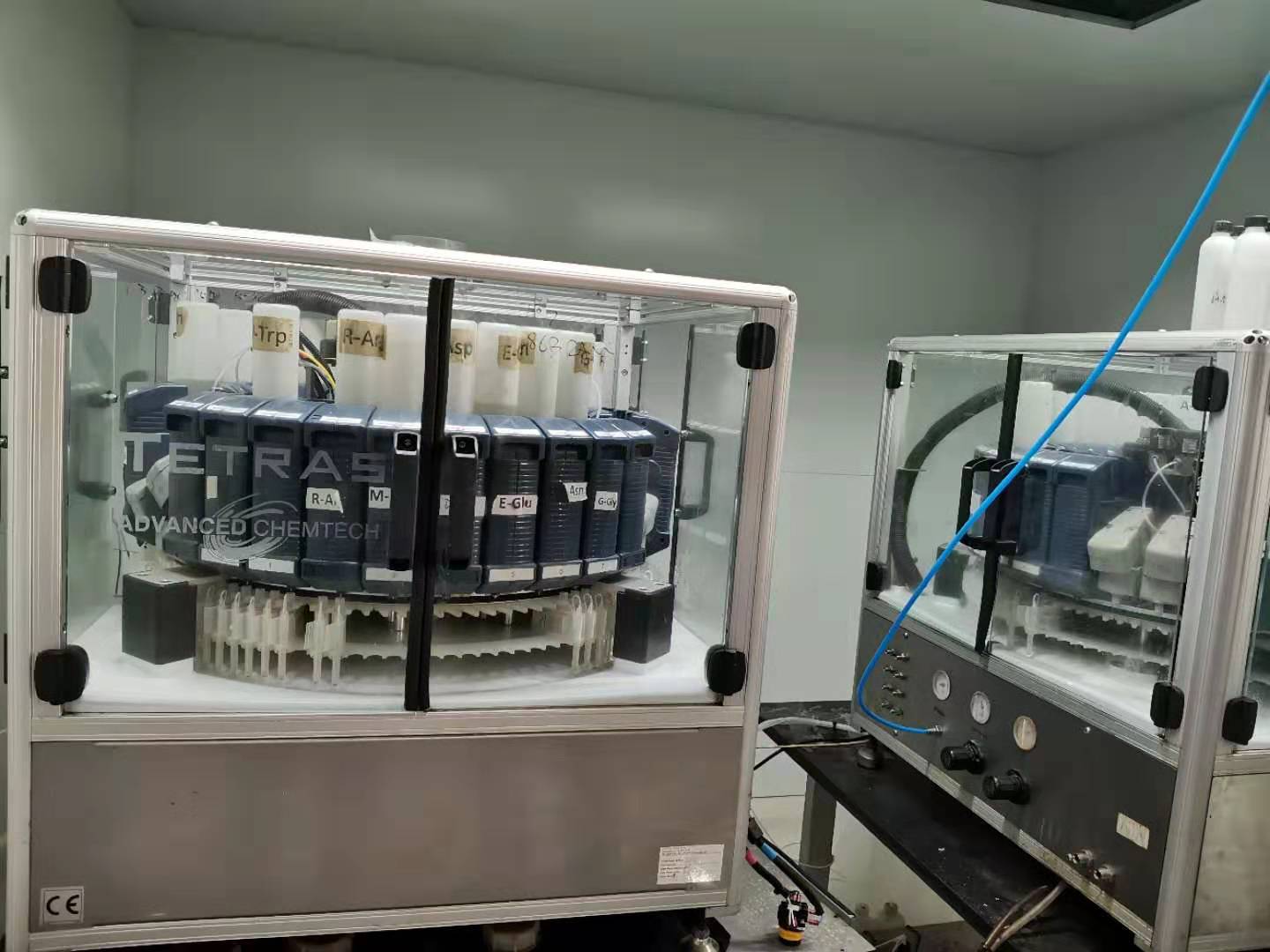

本公司的固相多肽合成技术具有如下特点:
与原有技术相比较,提高了合成精度,缩短合成时间,使单个肽链的合成长度突破了100个氨基酸残基。
操作简单,能快速开发工艺;无需分离中间体,自动化程度高。
实现了高通量,能高效率地合成大容量肽库。
可选择性强,能普遍应用于任意的氨基酸组合和千变万化的特殊修饰。
庞大的生产规模,同时合成上千条不同序列的订单。
证书



欢迎随时与我们联系!
风险提示:丁香通仅作为第三方平台,为商家信息发布提供平台空间。用户咨询产品时请注意保护个人信息及财产安全,合理判断,谨慎选购商品,商家和用户对交易行为负责。对于医疗器械类产品,请先查证核实企业经营资质和医疗器械产品注册证情况。
 用户评价
用户评价 暂无用户评价
暂无用户评价 文献和实验
文献和实验- Structure and regulation of the chromatin remodeller ISWI, Nature, 2016, 540:466–469.
- Structural basis for recognition of an endogenous peptide by the plant receptor kinase PEPR1, Cell Research, 2015, 25:110–120.
一:公司介绍 武汉明皓生物科技股份有限公司(http://www.moonbiochem.com)坐落于武汉东湖高新技术区光谷生物城, 是由一批在生命科学领域具有相当丰富经验的归国博士及经验丰富的专家共同创办的专业的生物公司。武汉明皓生物始终以“一切以客户为本”为理念,致力于为从事生命科学研究的科研人员提供高质量的化合物,多肽,蛋白及抗体产品以及高效优质的生物技术外包服务。 武汉明皓生物科技股份有限公司重在以高品质高质量为基础,以高满意度的客户服务为立命之本。我们坚信我们良好的发展
采用模型系统进行研究,可以得到对蛋白质的立体化学有贡献的实质性理解。合成多肽为确定与蛋白质的结构和功能有关的构象及相互作用,提供了颇具吸引力的途径。 螺旋结构是蛋白质、核酸、多糖及许多合成多肽的共同结构特征。研究模型多肽在固态和溶液中的螺旋结构,对明确解释在纤维蛋白和球蛋白的构象研究中得出的结论具有重要意义。 多肽是由α氨基酸按下面的通式组成的聚酰胺,其中R是α氨基酸中的侧链: 根据下面的基本假定,Pauling和Corey设想α-螺旋是折叠多肽比较稳定的二级结构之一: 多肽残基是反式共
保存: 大多肽在-20℃很稳定,特别是冷冻干燥并保存在干燥器中,在将它们暴露于空气之前, 冷冻干燥多肽可以放于室温。这将是湿度影响减少,当无法冷冻干燥时,最好的方法是以小的工作样量存放。 对于含Cys, Met orTrP的多肽,脱氧缓冲剂对其溶解必不可少,因为这种多肽可易空气氧化, 在封瓶前,慢慢流过多肽的氮气或氩气也会降低氧化作用。含Gln或Asn的多肽也容易降解,所有这些肽与不含这些有问题解苷的那些肽相比,生命期有限。 溶解性: 大多肽的道选溶剂是超纯