The unique features of PiggyBac transposons are that there is no cargo limit and it is also reversible. Genomes containing an inserted PiggyBac vector can be transiently transfected with the Excision only PB transposase (cat# PB220PA-1) expression vector to remove the transposons from the genome, footprint-free.
PB-EF1α-MCS-IRES-Neo PiggyBac cDNA Cloning and Expression Vector

Easily deliver your gene-of-interest using PiggyBac – this vector uses the EF1α promoter to co-expresses your cDNA and a neomycin marker via an IRES element
DESCRIPTION
SIZE
CATALOG NUMBER
PRICE
QUANTITY
ADD TO CART
PB-EF1α-MCS-IRES-Neo cDNA cloning and expression vector
10 µg
PB533A-2
Please Call
Contact Us
Overview
Get easy, consistent transgenesis
Consistent and easy-to-use, SBI’s PiggyBac Transposon System includes cloning and expression vectors that come with a range of markers as well as both constitutive and inducible promoters. The PB-EF1α-MCS-IRES-Neo PiggyBac cDNA Cloning and Expression Vector (Cat.# PB533A-2) drives co-expression of your gene-of-interest and the neomycin marker from the moderate EF1α promoter. Co-expression is mediated by an IRES element upstream of the neomycin resistance gene.

Why use the PiggyBac Transposon System?
Easy, consistent transgenesis with no limits on cargo size—For transgenesis that’s easy, consistent, and not limited by cargo size, SBI’s PiggyBac Transposon System is an excellent choice. The system consists of a PiggyBac Vector and the Super PiggyBac Transposase which recognizes transposon-specific inverted terminal repeats (ITRs) and efficiently integrates the ITRs and intervening DNA into the genome at TTAA sites. The Super PiggyBac Transposase is delivered to the cell via the Super PiggyBac Transposase Expression Vector, which is co-transfected with one or more PiggyBac Vectors.
Footprint-free removal that leaves no PiggyBac sequences behind—In addition to ease-of-use, consistency, and the lack of limits on DNA insert size, what sets this system apart is the ability to reverse the integration reaction in a footprint-free way—with the Excision Only PiggyBac Transposase (Cat.# PB220PA-1), the ITRs and cargo that the Super PiggyBac Transposase integrates into the genome can be removed, leaving behind the original genomic sequence and nothing else.
- Make transgenic cell lines with a single transfection
- Integrate multiple PiggyBac Vectors in a single transfection
- Insert an expression cassette into human, mouse, and rat cells
- Deliver virtually any-sized DNA insert, from 10 – 100 kb
- Choose from PiggyBac Vectors that express your gene-of-interest from constitutive or inducible promoters and include a variety of markers
- Determine the number of integration events with the PiggyBac qPCR Copy Number Kit (# PBC100A-1)
Customer Agreements
Academic customers can purchase PiggyBac Transposon System components for internal research purposes for indefinite use, whereas commercial customers must sign a customer agreement for a six-month, limited-use license to test the technology.
For end user license information, see the following:
* SBI is fully licensed to distribute PiggyBac vectors as a partnership with Transposagen Biopharmaceuticals, Inc.
How It Works
The PiggyBac Transposon System’s Cut-and-Paste Mechanism
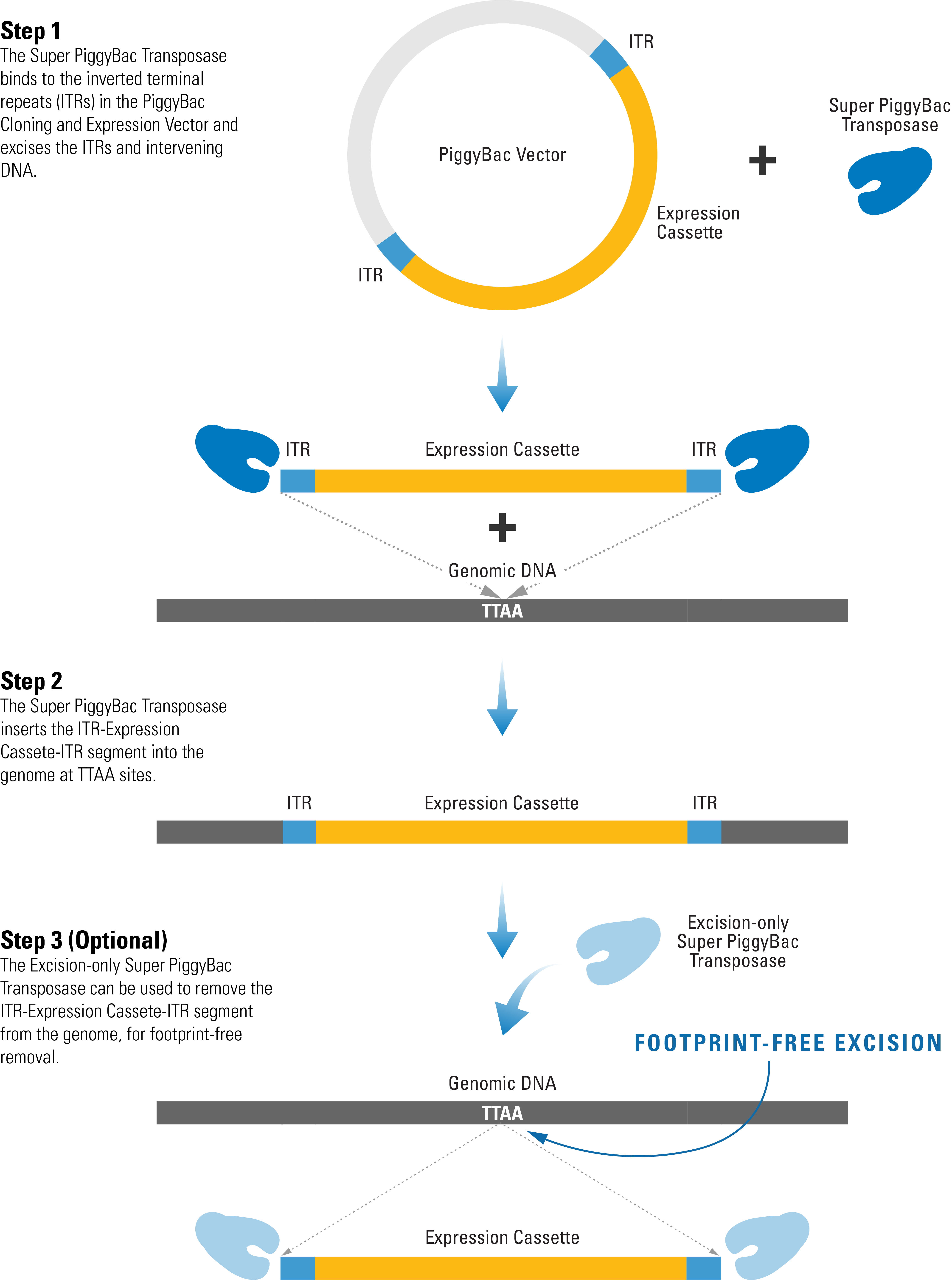
The efficient PiggyBac Transposon System uses a cut-and-paste mechanism to transfer DNA from the PiggyBac Vector into the genome. If only temporary genomic integration is desired, the Excision-only PiggyBac Transposase can be transiently expressed for footprint-free removal of the insert, resulting in reconstitution of the original genome sequence.
Figure 1. The PiggyBac Transposon System’s cut-and-paste mechanism.
- The Super PiggyBac Transposase binds to specific inverted terminal repeats (ITRs) in the PiggyBac Cloning and Expression Vector and excises the ITRs and intervening DNA.
- The Super PiggyBac Transposase inserts the ITR-Expression Cassette-ITR segment into the genome at TTAA sites.
- The Excision-only Super PiggyBac Transposase can be used to remove the ITR-Expression Cassette-ITR segment from the genome, for footprint-free removal
Supporting Data
One transfection can integrate one or more genes that can be precisely removed
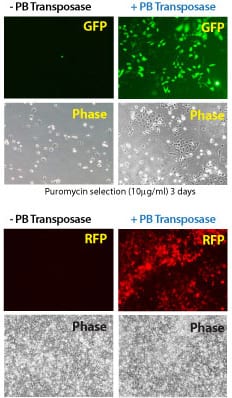
Figure 2. Efficient transgenesis with the Super PiggyBac Transposase and both single- and dual-promoter PiggyBac Vectors. (Top four panels) Co-transfection with the Super PiggyBac Transposase Expression Vector (Cat.# PB210PA-1) and a Dual Promoter PiggyBac Cloning and Expression Vector (Cat.# PB513B-1) into HeLa cells demonstrates the efficient integration delivered by SBI’s PiggyBac Transposon System. After ten days of puromycin selection, only the cells co-transfected with the Super PiggyBac Transposase (+PB, right two panels) show robust growth and GFP fluorescence. (Bottom four panels) Co-transfection with the Super PiggyBac Transposase Expression Vector (Cat.# PB210PA-1) and a Single Promoter PiggyBac Cloning and Expression Vector (Cat.# PB531A-2) into HEK293 cells further demonstrates the efficient integration delivered by SBI’s PiggyBac Transposon System. After seven days of growth, the majority of cells that received the Super PiggyBac Transposase Expression Vector (+PB, right two panels) were RFP positive.
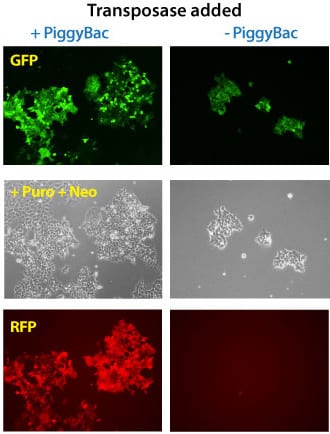
Figure 3. Simultaneous integration of multiple PiggyBac Vectors is also highly efficient. METHODS: Three different PiggyBac transposon vectors (Cat.# PB513B-1, Cat.# PB533A-2, and Cat.# PB531A-2) were co-transfected with (left panels) or without (right panels) the Super PiggyBac Transposase Expression Vector (Cat.# PB210PA-1) into Human HT1080 cells. Puromycin and neomycin selection was applied for seven days. The cells that were co-transfected with the Super PiggyBac Transposase Expression Vector were puro and neo resistant, GFP-positive, and RFP-positive. Background GFP-positive cells that are puro resistant stem from random PB513B-1 integrations during the puromycin selection. The non-PiggyBac-mediated integration rate in those cells was extremely low and no RFP-positive cells were identified.
Product Documentation
Citations
- Shinmura, K, et al. (2017) WDR62 overexpression is associated with a poor prognosis in patients with lung adenocarcinoma. Mol. Carcinog.. 2017 Aug 1; 56(8):1984-1991. PM ID: 28277612
- Henssen, AG, et al. (2017) PGBD5 promotes site-specific oncogenic mutations in human tumors. Nat. Genet.. 2017 Jul 1; 49(7):1005-1014. PM ID:28504702
- Katayama, M, et al. (2017) Immortalized prairie vole-derived fibroblasts (VMF-K4DTs) can be transformed into pluripotent stem cells and provide a useful tool with which to determine optimal reprogramming conditions. J. Reprod. Dev.. 2017 Jun 21; 63(3):311-318. PM ID: 28331164
- Cammareri, P, et al. (2017) TGFβ pathway limits dedifferentiation following WNT and MAPK pathway activation to suppress intestinal tumourigenesis. Cell Death Differ.. 2017 Jun 16;. PM ID: 28622298
- Kimura, T, et al. (2017) Hsp90 inhibitor geldanamycin attenuates the cytotoxicity of sunitinib in cardiomyocytes via inhibition of the autophagy pathway. Toxicol. Appl. Pharmacol.. 2017 Jun 15; 329:282-292. PM ID: 28624441
- Maegawa, KI, et al. (2017) The Highly Dynamic Nature of ERdj5 Is Key to Efficient Elimination of Aberrant Protein Oligomers through ER-Associated Degradation. Structure. 2017 Jun 6; 25(6):846-857.e4. PM ID: 28479060
- Liu, H, et al. (2017) Stochastic Protein Labeling Enables Long-Term Single Molecule Observation in vivo. bioRxiv. 2017 Jun 5;. Link: bioRxiv
- Park, TS, Kim, SW & Lee, JH. (2017) Efficient transgene expression system using a cumate-inducible promoter and Cre-loxP recombination in avian cells. Asian-australas. J. Anim. Sci.. 2017 Jun 1; 30(6):886-892. PM ID: 27764912
- Lin, W & Li, Z. (2017) Blueberries inhibit cyclooxygenase-1 and cyclooxygenase-2 activity in human epithelial ovarian cancer. Oncol Lett. 2017 Jun 1; 13(6):4897-4904. PM ID: 28599493
- Yamazaki, T, et al. (2017) Targeted DNA methylation in pericentromeres with genome editing-based artificial DNA methyltransferase. PLoS ONE. 2017 May 25; 12(5):e0177764. PM ID: 28542388
- Herrington, KA, et al. (2017) Spatial analysis of Cdc42 activity reveals a role for plasma membrane-associated Cdc42 in centrosome regulation. Mol. Biol. Cell. 2017 May 24;. PM ID: 28539409
- Cruz-Molina, S, et al. (2017) PRC2 Facilitates the Regulatory Topology Required for Poised Enhancer Function during Pluripotent Stem Cell Differentiation. Cell Stem Cell. 2017 May 4; 20(5):689-705.e9. PM ID: 28285903
- Wang, L, et al. (2017) Derivation and characterization of primordial germ cells from Guangxi yellow-feather chickens.. Poult. Sci.. 2017 May 1; 96(5):1419-1425. PM ID: 28158811
- Uehara, T, et al. (2017) Selective degradation of splicing factor CAPER [alpha] by anticancer sulfonamides. Nature Chemical Biology. 2017 Apr 27; 13:675–680. Link: Nature Chemical Biology
- Qi, Z, et al. (2017) An optimized, broadly applicable piggyBac transposon induction system. Nucleic Acids Res.. 2017 Apr 20; 45(7):e55. PM ID: 28082389
- Wen, Y, et al. (2017) A stable but reversible integrated surrogate reporter for assaying CRISPR/Cas9-stimulated homology-directed repair. J. Biol. Chem.. 2017 Apr 14; 292(15):6148-6162. PM ID: 28228480
- Mitra, A, et al. (2017) IL6-mediated inflammatory loop reprograms normal to epithelial-mesenchymal transition(+) metastatic cancer stem cells in preneoplastic liver of transforming growth factor beta-deficient β2-spectrin(+/-) mice. Hepatology. 2017 Apr 1; 65(4):1222-1236. PM ID: 27863449
- Zamboni, CG, et al. (2017) Polymeric nanoparticles as cancer-specific DNA delivery vectors to human hepatocellular carcinoma. J Control Release. 2017 Mar 27;. PM ID: 28351668
- Katayama, H, et al. (2017) Generation of non-viral, transgene-free hepatocyte like cells with piggyBac transposon. Sci Rep. 2017 Mar 15; 7:44498. PM ID:28295042
- Xu, X, et al. (2017) Reversal of Phenotypic Abnormalities by CRISPR/Cas9-Mediated Gene Correction in Huntington Disease Patient-Derived Induced Pluripotent Stem Cells.. Stem Cell Reports. 2017 Mar 14; 8(3):619-633. PM ID: 28238795






 文献和实验
文献和实验


