相关产品推荐更多 >
万千商家帮你免费找货
0 人在求购买到急需产品
- 详细信息
- 文献和实验
- 技术资料
- 英文名:
HCC95
- 库存:
100
- 供应商:
中乔新舟
- 品系:
细胞系
- 运输方式:
常温
- 年限:
液氮长期
- 生长状态:
贴壁生长
- 规格:
T25
|
产品名称 |
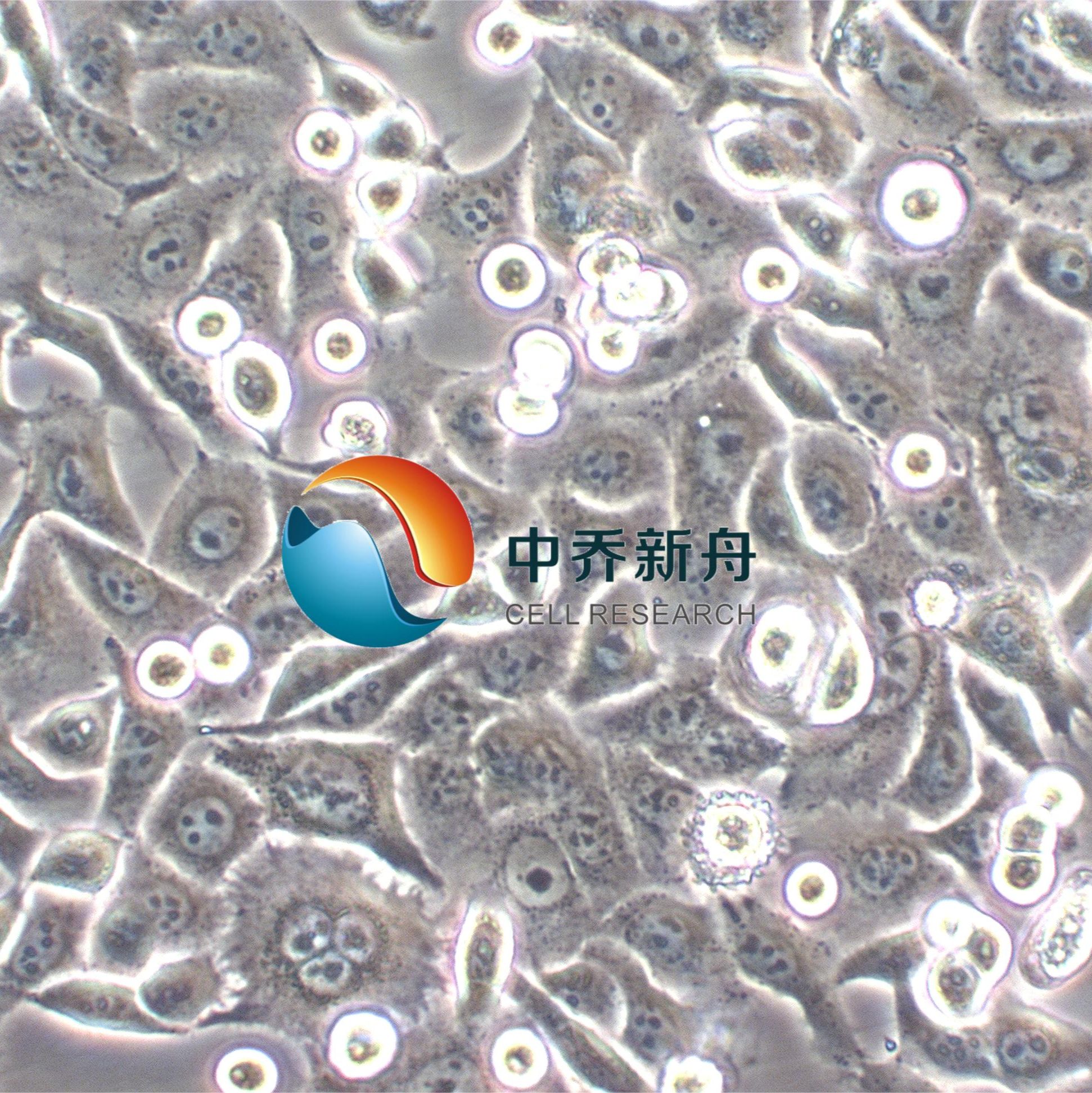
HCC95人肺鳞癌细胞 |
|
货号 |
ZQ0978 |
|
产品介绍 |
HCC95细胞系源自一位65岁男性的肺鳞状细胞癌组织,细胞表现出上皮细胞样的形态,并且是贴壁生长。主要应用于药物筛选和敏感性测试、癌症生物学研究、癌症基因组学和蛋白质组学研究、癌症转移研究、免疫治疗研究、耐药性研究、癌症治疗靶点的验证等研究。 |
|
种属 |
人 |
|
性别/年龄 |
男性/65岁 |
|
组织 |
肺 |
|
疾病 |
鳞状细胞癌 |
|
生物安全等级 |
BSL-1 |
|
STR位点信息 |
D3S1358: 17 |
|
细胞类型 |
肿瘤细胞 |
|
形态学 |
上皮 |
|
生长方式 |
贴壁 |
|
倍增时间 |
40.6 hours (PubMed=29681454) |
|
培养基和添加剂 |
RPMI-1640(中乔新舟 货号:ZQ-200)+10%FBS(品牌:中乔新舟 货号:ZQ0500)+1%PS(中乔新舟 货号:CSP006) |
|
推荐完全培养基货号 |
ZM0978 |
|
培养条件 |
95%空气,5%二氧化碳;37℃ |
|
抗原表达/受体表达 |
*** |
|
基因表达 |
*** |
|
保藏机构 |
KCLB; 70095 |
|
供应限制 |
仅供科研使用 |
上海中乔新舟生物科技有限公司成立于2011年,历经十多年发展,主要专注于细胞生物学产品的研究和开发,专注于为药企、各类科研机构及CRO企业提供符合标准规范的细胞培养服务、细胞培养基、细胞检测试剂盒、细胞培养试剂,胎牛血清和细胞生物学技术服务等。
公司一直致力于为高等院校、研究机构、医院、CRO及CDMO企业提供细胞培养完整解决方案,这些产品旨在满足细胞培养的多样需求,确保实验和研究的有效进行。引用中乔新舟(ZQXZBIO)产品和服务的文献超数千篇。

产品服务
细胞资源:原代细胞、细胞株、干细胞、示踪细胞、耐药株细胞、永生化细胞等基因工程细胞。
试剂产品:胎牛血清、完全培养基(适用于原代细胞及细胞株)、无血清培养基、基础培养基、细胞转染试剂、重组因子、胰酶和双抗等等细胞培养所有实验相关产品。
技术服务:稳转株构建、原代细胞分离、特殊培养基定制服务、细胞检测等。

目前产品已经畅销国内30多个省市,与客户建立长期的合作伙伴关系,共同实现成功。全体员工将不懈努力,继续为科研人员提供优良的产品和服务,致力成为全球细胞培养领域的参与者。
企业愿景
致力于成为国内细胞培养基产业的佼佼者,生物医药领域上游原材料的优良提供商。
企业使命
成长为专业细胞系及原代细胞培养供应商、专业细胞培养基及培养试剂生产商。
企业荣誉

风险提示:丁香通仅作为第三方平台,为商家信息发布提供平台空间。用户咨询产品时请注意保护个人信息及财产安全,合理判断,谨慎选购商品,商家和用户对交易行为负责。对于医疗器械类产品,请先查证核实企业经营资质和医疗器械产品注册证情况。
 文献和实验
文献和实验
PubMed=9559342; DOI=10.1002/(SICI)1098-2264(199804)21:4<308::AID-GCC4>3.0.CO;2-2
Virmani A.K., Fong K.M., Kodagoda D.R., McIntire D., Hung J., Tonk V., Minna J.D., Gazdar A.F.
Allelotyping demonstrates common and distinct patterns of chromosomal loss in human lung cancer types.
Genes Chromosomes Cancer 21:308-319(1998)
PubMed=10353731
Wistuba I.I., Bryant D., Behrens C., Milchgrub S., Virmani A.K., Ashfaq R., Minna J.D., Gazdar A.F.
Comparison of features of human lung cancer cell lines and their corresponding tumors.
Clin. Cancer Res. 5:991-1000(1999)
PubMed=10987304
Girard L., Zochbauer-Muller S., Virmani A.K., Gazdar A.F., Minna J.D.
Genome-wide allelotyping of lung cancer identifies new regions of allelic loss, differences between small cell lung cancer and non-small cell lung cancer, and loci clustering.
Cancer Res. 60:4894-4906(2000)
PubMed=16187286; DOI=10.1002/ijc.21491
Garnis C., Lockwood W.W., Vucic E., Ge Y., Girard L., Minna J.D., Gazdar A.F., Lam S., MacAulay C., Lam W.L.
High resolution analysis of non-small cell lung cancer cell lines by whole genome tiling path array CGH.
Int. J. Cancer 118:1556-1564(2006)
PubMed=19472407; DOI=10.1002/humu.21028
Blanco R., Iwakawa R., Tang M.-Y., Kohno T., Angulo B., Pio R., Montuenga L.M., Minna J.D., Yokota J., Sanchez-Cespedes M.
A gene-alteration profile of human lung cancer cell lines.
Hum. Mutat. 30:1199-1206(2009)
PubMed=20557307; DOI=10.1111/j.1349-7006.2010.01622.x
Iwakawa R., Kohno T., Enari M., Kiyono T., Yokota J.
Prevalence of human papillomavirus 16/18/33 infection and p53 mutation in lung adenocarcinoma.
Cancer Sci. 101:1891-1896(2010)
PubMed=20679594; DOI=10.1093/jnci/djq279
Gazdar A.F., Girard L., Lockwood W.W., Lam W.L., Minna J.D.
Lung cancer cell lines as tools for biomedical discovery and research.
J. Natl. Cancer Inst. 102:1310-1321(2010)
PubMed=22460905; DOI=10.1038/nature11003
Barretina J.G., Caponigro G., Stransky N., Venkatesan K., Margolin A.A., Kim S., Wilson C.J., Lehar J., Kryukov G.V., Sonkin D., Reddy A., Liu M., Murray L., Berger M.F., Monahan J.E., Morais P., Meltzer J., Korejwa A., Jane-Valbuena J., Mapa F.A., Thibault J., Bric-Furlong E., Raman P., Shipway A., Engels I.H., Cheng J., Yu G.-Y.K., Yu J.-J., Aspesi P. Jr., de Silva M., Jagtap K., Jones M.D., Wang L., Hatton C., Palescandolo E., Gupta S., Mahan S., Sougnez C., Onofrio R.C., Liefeld T., MacConaill L.E., Winckler W., Reich M., Li N.-X., Mesirov J.P., Gabriel S.B., Getz G., Ardlie K., Chan V., Myer V.E., Weber B.L., Porter J., Warmuth M., Finan P., Harris J.L., Meyerson M.L., Golub T.R., Morrissey M.P., Sellers W.R., Schlegel R., Garraway L.A.
The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity.
Nature 483:603-607(2012)
PubMed=22961666; DOI=10.1158/2159-8290.CD-12-0112
Byers L.A., Wang J., Nilsson M.B., Fujimoto J., Saintigny P., Yordy J., Giri U., Peyton M., Fan Y.-H., Diao L.-X., Masrorpour F., Shen L., Liu W.-B., Duchemann B., Tumula P., Bhardwaj V., Welsh J., Weber S., Glisson B.S., Kalhor N., Wistuba I.I., Girard L., Lippman S.M., Mills G.B., Coombes K.R., Weinstein J.N., Minna J.D., Heymach J.V.
Proteomic profiling identifies dysregulated pathways in small cell lung cancer and novel therapeutic targets including PARP1.
Cancer Discov. 2:798-811(2012)
PubMed=24002593; DOI=10.1038/bjc.2013.452
Wu D., Pang Y., Wilkerson M.D., Wang D., Hammerman P.S., Liu J.S.
Gene-expression data integration to squamous cell lung cancer subtypes reveals drug sensitivity.
Br. J. Cancer 109:1599-1608(2013)
PubMed=25877200; DOI=10.1038/nature14397
Yu M., Selvaraj S.K., Liang-Chu M.M.Y., Aghajani S., Busse M., Yuan J., Lee G., Peale F.V., Klijn C., Bourgon R., Kaminker J.S., Neve R.M.
A resource for cell line authentication, annotation and quality control.
Nature 520:307-311(2015)
PubMed=26589293; DOI=10.1186/s13073-015-0240-5
Scholtalbers J., Boegel S., Bukur T., Byl M., Goerges S., Sorn P., Loewer M., Sahin U., Castle J.C.
TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression.
Genome Med. 7:118.1-118.7(2015)
PubMed=26361996; DOI=10.1016/j.jprot.2015.09.003
Grundner-Culemann K., Dybowski J.N., Klammer M., Tebbe A., Schaab C., Daub H.
Comparative proteome analysis across non-small cell lung cancer cell lines.
J. Proteomics 130:1-10(2016)
PubMed=28196595; DOI=10.1016/j.ccell.2017.01.005
Li J., Zhao W., Akbani R., Liu W.-B., Ju Z.-L., Ling S.-Y., Vellano C.P., Roebuck P., Yu Q.-H., Eterovic A.K., Byers L.A., Davies M.A., Deng W.-L., Gopal Y.N.V., Chen G., von Euw E.M., Slamon D.J., Conklin D., Heymach J.V., Gazdar A.F., Minna J.D., Myers J.N., Lu Y.-L., Mills G.B., Liang H.
Characterization of human cancer cell lines by reverse-phase protein arrays.
Cancer Cell 31:225-239(2017)
PubMed=29610121; DOI=10.1158/2159-8290.CD-17-1004
Drilon A., Somwar R., Mangatt B.P., Edgren H., Desmeules P., Ruusulehto A., Smith R.S., Delasos L., Vojnic M., Plodkowski A.J., Sabari J., Ng K., Montecalvo J., Chang J., Tai H.-C., Lockwood W.W., Martinez V., Riely G.J., Rudin C.M., Kris M.G., Arcila M.E., Matheny C., Benayed R., Rekhtman N., Ladanyi M., Ganji G.
Response to ERBB3-directed targeted therapy in NRG1-rearranged cancers.
Cancer Discov. 8:686-695(2018)
PubMed=29681454; DOI=10.1016/j.cell.2018.03.028
McMillan E.A., Ryu M.-J., Diep C.H., Mendiratta S., Clemenceau J.R., Vaden R.M., Kim J.-H., Motoyaji T., Covington K.R., Peyton M., Huffman K., Wu X.-F., Girard L., Sung Y., Chen P.-H., Mallipeddi P.L., Lee J.Y., Hanson J., Voruganti S., Yu Y., Park S., Sudderth J., DeSevo C., Muzny D.M., Doddapaneni H., Gazdar A.F., Gibbs R.A., Hwang T.H., Heymach J.V., Wistuba I.I., Coombes K.R., Williams N.S., Wheeler D.A., MacMillan J.B., DeBerardinis R.J., Roth M.G., Posner B.A., Minna J.D., Kim H.S., White M.A.
Chemistry-first approach for nomination of personalized treatment in lung cancer.
Cell 173:864-878.e29(2018)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=31068700; DOI=10.1038/s41586-019-1186-3
Ghandi M., Huang F.W., Jane-Valbuena J., Kryukov G.V., Lo C.C., McDonald E.R. III, Barretina J.G., Gelfand E.T., Bielski C.M., Li H.-X., Hu K., Andreev-Drakhlin A.Y., Kim J., Hess J.M., Haas B.J., Aguet F., Weir B.A., Rothberg M.V., Paolella B.R., Lawrence M.S., Akbani R., Lu Y.-L., Tiv H.L., Gokhale P.C., de Weck A., Mansour A.A., Oh C., Shih J., Hadi K., Rosen Y., Bistline J., Venkatesan K., Reddy A., Sonkin D., Liu M., Lehar J., Korn J.M., Porter D.A., Jones M.D., Golji J., Caponigro G., Taylor J.E., Dunning C.M., Creech A.L., Warren A.C., McFarland J.M., Zamanighomi M., Kauffmann A., Stransky N., Imielinski M., Maruvka Y.E., Cherniack A.D., Tsherniak A., Vazquez F., Jaffe J.D., Lane A.A., Weinstock D.M., Johannessen C.M., Morrissey M.P., Stegmeier F., Schlegel R., Hahn W.C., Getz G., Mills G.B., Boehm J.S., Golub T.R., Garraway L.A., Sellers W.R.
Next-generation characterization of the Cancer Cell Line Encyclopedia.
Nature 569:503-508(2019)
PubMed=31395879; DOI=10.1038/s41467-019-11415-2
Yu K., Chen B., Aran D., Charalel J., Yau C., Wolf D.M., van 't Veer L.J., Butte A.J., Goldstein T., Sirota M.
Comprehensive transcriptomic analysis of cell lines as models of primary tumors across 22 tumor types.
Nat. Commun. 10:3574.1-3574.11(2019)
PubMed=31803961; DOI=10.1002/jcb.29564
Mulshine J.L., Ujhazy P., Antman M., Burgess C.M., Kuzmin I.A., Bunn P.A. Jr., Johnson B.E., Roth J.A., Pass H.I., Ross S.M., Aldige C.R., Wistuba I.I., Minna J.D.
From clinical specimens to human cancer preclinical models -- a journey the NCI-cell line database-25 years later.
J. Cell. Biochem. 121:3986-3999(2020)
PubMed=31978347; DOI=10.1016/j.cell.2019.12.023
Nusinow D.P., Szpyt J., Ghandi M., Rose C.M., McDonald E.R. III, Kalocsay M., Jane-Valbuena J., Gelfand E.T., Schweppe D.K., Jedrychowski M.P., Golji J., Porter D.A., Rejtar T., Wang Y.K., Kryukov G.V., Stegmeier F., Erickson B.K., Garraway L.A., Sellers W.R., Gygi S.P.
Quantitative proteomics of the Cancer Cell Line Encyclopedia.
Cell 180:387-402.e16(2020)
双向凝胶电泳-飞行时间质谱法分析人肺鳞癌细胞HCL-H520蛋白质组成份
《双向凝胶电泳-飞行时间质谱法分析人肺鳞癌细胞HCL-H520蛋白质组成份》下载 点击 这里 下载
三句话读懂一篇 CNS:让人产生幻觉的毒蘑菇,竟能治疗抑郁症?韩春雨开发 RNA 高效追踪平台
:接种新冠加强针更能抵抗奥密克戎 2022 年 1 月 28 日,中国科学院生物物理研究所王祥喜等多个团队联合在 Nature 杂志发表研究论文 Memory B cell repertoire from triple vaccinees against diverse SARS-CoV-2 variants。 该研究检查了接受两剂或三剂灭活疫苗的个体的血清是否可以中和真正的 Omicron,结果显示 2 剂和 3 剂疫苗的中和抗体的血清转化率分别为 3.3%(2/60)和 95%(57
骨髓瘤细胞 1000元 FRhK-4 GNO18 恒河猴 恒河猴胚肾细胞 900元 GBC-SD TCHu 16 人 胆囊癌细胞 400元 H9c2(2-1) GNR 5 大鼠 心肌细胞 1000元 HA GNHu 4 人 羊膜细胞 400元 HBL-100 GNHu10 人 整合SV40基因的乳腺上皮细胞 400元 HCC 94 [HCC941122] TCHu 81 人 子宫鳞癌细胞(高分化) 400元 HCCC-9810
 技术资料
技术资料暂无技术资料 索取技术资料




![293 [HEK-293]-RED人胚肾细胞-红色标记](https://img1.dxycdn.com/p/s14/2025/0402/220/4815421658941627091.jpg!wh200)