手机验证

SAS
详询
SAS
详询
无
博辉生物科技(广州)有限公司
1x10^6 cells
无
SAS
贴壁
不限
干冰/常温
人舌头鳞状肿瘤细胞
是
上皮细胞样
未知
人/Human
人舌头鳞状肿瘤细胞
人舌头鳞状肿瘤细胞
冻存管/培养瓶
| 目录号 | BH-C342 |
| 细胞英文(简称) | SAS |
| 细胞名称 | SAS;人舌头鳞状肿瘤细胞 |
| 背景资料 | 收到细胞,请查看瓶子是否有破裂,培养基是否漏出,是否浑浊,如有请尽快和我们联系。如包装完好,取出25cm2培养瓶,75%酒精消毒,拆下封口膜,在显微镜下观察细胞形态是否正常,细胞的贴壁情况以及细胞密度,然后放入37℃,5%CO2细胞培养箱中静置2-3小时,以稳定细胞状态。 |
| 细胞来源 | JCRB |
| 代次 | P3 |
| 规格 | 冻存管/培养瓶 |
| 细胞数 | 1x10^6 cells |
| 生物安全级别 | 1 |
| 组织来源 | 舌头 |
| 细胞形态 | 贴壁 |
| 细胞活力 | 95%(Viability by Trypan Blue Exclusion) |
| 细胞检测 | 细胞不含有HIV-1、HBV、HCV、支原体、细菌、酵母和真菌 |
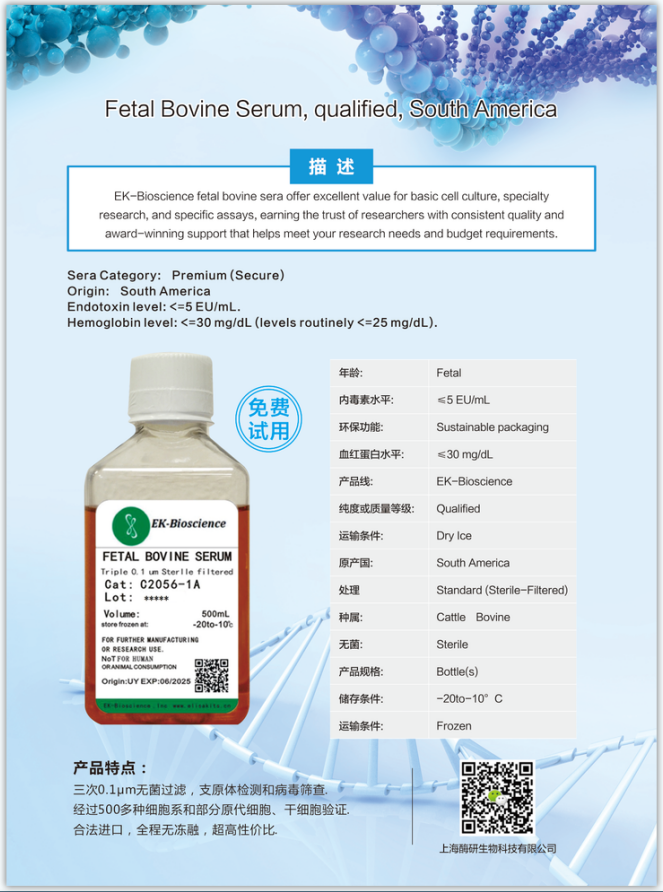
| 培养条件 | DMEM/F12+10%FBS(货号:C2056-1A)+1%P/S;37℃,5% CO2 |
| 传代方法 | 建议1:2-1:3 两天换液一次 |
| 冻存条件 | 无血清细胞冻存液(货号:S002) |




风险提示:丁香通仅作为第三方平台,为商家信息发布提供平台空间。用户咨询产品时请注意保护个人信息及财产安全,合理判断,谨慎选购商品,商家和用户对交易行为负责。对于医疗器械类产品,请先查证核实企业经营资质和医疗器械产品注册证情况。
资料下载:
SAS(BH-C342).docx 附件下载 (下载 0 次)
A5培养操作说明.pdf 附件下载 (下载 0 次)
A5细胞收到处理方法.pdf 附件下载 (下载 0 次)
请 [登录] 后再下载!
 文献和实验
文献和实验
DOI=10.11277/stomatology1952.38.20
Takahashi K., Kanazawa H., Akiyama Y., Tazaki S., Takahara M., Muto T., Tanzawa H., Sato K.-I.
Establishment and characterization of a cell line (SAS) from poorly differentiated human squamous cell carcinoma of the tongue.
J. Jpn. Stomatol. Soc. 38:20-28(1989)
PubMed=9023415; DOI=10.1006/cimm.1996.1062
Seki N., Hoshino T., Kikuchi M., Hayashi A., Itoh K.
HLA-A locus-restricted and tumor-specific CTLs in tumor-infiltrating lymphocytes of patients with non-small cell lung cancer.
Cell. Immunol. 175:101-110(1997)
PubMed=9178645; DOI=10.1006/cimm.1997.1108
Nakao M., Sata M., Saitsu H., Yutani S., Kawamoto M., Kojiro M., Itoh K.
CD4+ hepatic cancer-specific cytotoxic T lymphocytes in patients with hepatocellular carcinoma.
Cell. Immunol. 177:176-181(1997)
PubMed=9290701; DOI=10.1002/(SICI)1098-2744(199708)19:4<243::aid-mc5>3.0.CO;2-D
Jia L.-Q., Osada M., Ishioka C., Gamo M., Ikawa S., Suzuki T., Shimodaira H., Niitani T., Kudo T., Akiyama M., Kimura N., Matsuo M., Mizusawa H., Tanaka N., Koyama H., Namba M., Kanamaru R., Kuroki T.
Screening the p53 status of human cell lines using a yeast functional assay.
Mol. Carcinog. 19:243-253(1997)
DOI=10.5843/jsot.13.Suppliment_301
Tadateru A., Tarou I., Tachikawa T.
Establishment and charactalization of human oral squamous cell carcinoma cell line with highly lymph node metastatic potential.
J. Jpn. Soc. Oral Oncol. 13 Suppl. 1:301-306(2001)
PubMed=17325662; DOI=10.1038/sj.onc.1210330
Nakaya K., Yamagata H.D., Arita N., Nakashiro K.-I., Nose M., Miki T., Hamakawa H.
Identification of homozygous deletions of tumor suppressor gene FAT in oral cancer using CGH-array.
Oncogene 26:5300-5308(2007)
PubMed=20164919; DOI=10.1038/nature08768
Bignell G.R., Greenman C.D., Davies H., Butler A.P., Edkins S., Andrews J.M., Buck G., Chen L., Beare D., Latimer C., Widaa S., Hinton J., Fahey C., Fu B.-Y., Swamy S., Dalgliesh G.L., Teh B.T., Deloukas P., Yang F.-T., Campbell P.J., Futreal P.A., Stratton M.R.
Signatures of mutation and selection in the cancer genome.
Nature 463:893-898(2010)
PubMed=20215515; DOI=10.1158/0008-5472.CAN-09-3458
Rothenberg S.M., Mohapatra G., Rivera M.N., Winokur D., Greninger P., Nitta M., Sadow P.M., Sooriyakumar G., Brannigan B.W., Ulman M.J., Perera R.M., Wang R., Tam A., Ma X.-J., Erlander M., Sgroi D.C., Rocco J.W., Lingen M.W., Cohen E.E.W., Louis D.N., Settleman J., Haber D.A.
A genome-wide screen for microdeletions reveals disruption of polarity complex genes in diverse human cancers.
Cancer Res. 70:2158-2164(2010)
PubMed=24627082; DOI=10.3892/ijo.2014.2332
Fujinaga T., Kumamaru W., Sugiura T., Kobayashi Y., Ohyama Y., Ikari T., Onimaru M., Akimoto N., Jogo R., Mori Y.
Biological characterization and analysis of metastasis-related genes in cell lines derived from the primary lesion and lymph node metastasis of a squamous cell carcinoma arising in the mandibular gingiva.
Int. J. Oncol. 44:1614-1624(2014)
PubMed=27397505; DOI=10.1016/j.cell.2016.06.017
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., Aben N., Goncalves E., Barthorpe S., Lightfoot H., Cokelaer T., Greninger P., van Dyk E., Chang H., de Silva H., Heyn H., Deng X.-M., Egan R.K., Liu Q.-S., Mironenko T., Mitropoulos X., Richardson L., Wang J.-H., Zhang T.-H., Moran S., Sayols S., Soleimani M., Tamborero D., Lopez-Bigas N., Ross-Macdonald P., Esteller M., Gray N.S., Haber D.A., Stratton M.R., Benes C.H., Wessels L.F.A., Saez-Rodriguez J., McDermott U., Garnett M.J.
A landscape of pharmacogenomic interactions in cancer.
Cell 166:740-754(2016)
PubMed=30894373; DOI=10.1158/0008-5472.CAN-18-2747
Dutil J., Chen Z.-H., Monteiro A.N.A., Teer J.K., Eschrich S.A.
An interactive resource to probe genetic diversity and estimated ancestry in cancer cell lines.
Cancer Res. 79:1263-1273(2019)
PubMed=35839778; DOI=10.1016/j.ccell.2022.06.010
Goncalves E., Poulos R.C., Cai Z.-X., Barthorpe S., Manda S.S., Lucas N., Beck A., Bucio-Noble D., Dausmann M., Hall C., Hecker M., Koh J., Lightfoot H., Mahboob S., Mali I., Morris J., Richardson L., Seneviratne A.J., Shepherd R., Sykes E., Thomas F., Valentini S., Williams S.G., Wu Y.-X., Xavier D., MacKenzie K.L., Hains P.G., Tully B., Robinson P.J., Zhong Q., Garnett M.J., Reddel R.R.
Pan-cancer proteomic map of 949 human cell lines.
Cancer Cell 40:835-849.e8(2022)
 技术资料
技术资料


